|
|
|
|
|
|
|
|
|
Award Number and Duration |
|
|
|
NSF DBI 1661375 |
|
|
|
Point of Contact and PI |
|
|
|
Bei Wang (PI, University of Utah) |
|
|
|
Collaborators |
|
|
|
Anantharaman Kalyanaraman (PI, Washington State University) |
|
|
|
Overview |
|
|
|
Understanding how gene by environment interactions result in specific phenotypes is a core goal of modern biology and has real-world impacts on such things as crop management. Developing and managing successful crop practices is a goal that is fundamentally tied to our national food security. By applying novel computational visual analytical methods, this project seeks to identify and unravel the complex web of interactions linking genotypes, environments and phenotypes. These methods will first need to be designed and developed into usable software applications that can handle large volumes of crop phenomics data. High-throughput sensing technologies collect large volumes of field data for many plant traits, such as flowering time, related to crop development and production. The maize cultivars used here come from multiple genotypes that have been grown under a variety of environmental conditions, in order to give the widest range of conditions for understanding the interactions. The resulting data sets are growing quickly, both in size and complexity, but the analytical tools needed to extract knowledge and catalyze scientific discoveries have significantly lagged behind. The methodologies to be developed in this project represent a systematic attempt at bridging this rapidly widening divide. The project is inherently interdisciplinary, involving close research partnerships among computer scientists, plant scientists, and mathematicians. The research outcomes will be tightly integrated with education using a multipronged approach that includes, among others, postdoctoral and student training (graduates and undergraduates), curriculum development for a new campus-wide interdisciplinary undergraduate degree in Data Analytics, conference tutorials for training phenomics data practitioners, and contribution to the recruitment and retention of underrepresented minorities (particularly women) in STEM fields through the Pacific Northwest Louis Stokes Alliance for Minority Participation. |
|
|
|
Publications |
|
|
| Year 4 (2020 - 2021) | |
|
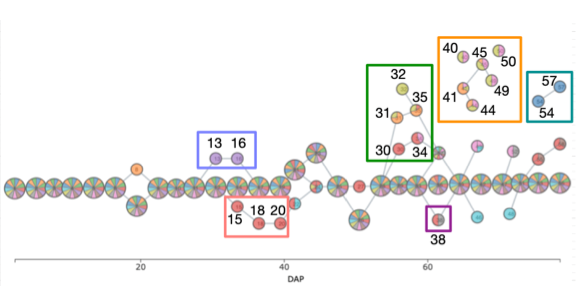
 Pheno-Mapper: An Interactive Toolbox for the Visual Exploration of Phenomics Data.
Pheno-Mapper: An Interactive Toolbox for the Visual Exploration of Phenomics Data.
Youjia Zhou, Methun Kamruzzaman, Patrick Schnable, Bala Krishnamoorthy, Ananth Kalyanaraman, Bei Wang. Proceedings of the 12th ACM Conference on Bioinformatics, Computational Biology, and Health Informatics (ACM BCB), Article No. 20, pages 1-10, 2021. DOI: 10.1145/3459930.3469511 |

|
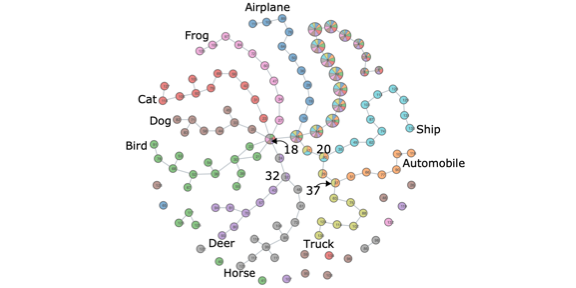
 Mapper Interactive: A Scalable, Extendable, and Interactive Toolbox for the Visual Exploration of High-Dimensional Data.
Mapper Interactive: A Scalable, Extendable, and Interactive Toolbox for the Visual Exploration of High-Dimensional Data.Youjia Zhou, Nithin Chalapathi, Archit Rathore, Yaodong Zhao, Bei Wang. IEEE Pacific Visualization Symposium, 2021. DOI: 10.1109/PacificVis52677.2021.00021 arXiv:2011.03209. |
|
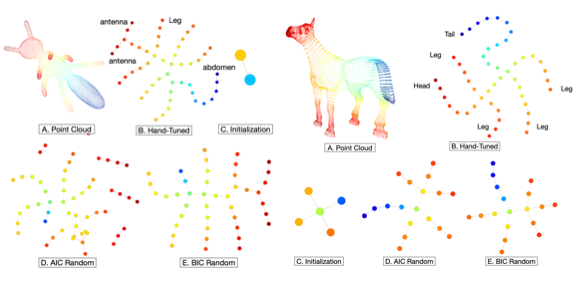
 Adaptive Covers for Mapper Graphs Using Information Criteria.
Adaptive Covers for Mapper Graphs Using Information Criteria.Nithin Chalapathi, Youjia Zhou, Bei Wang. Workshop on Applications of Topological Data Analysis to Big Data at the IEEE International Conference on Big Data, 2021. |
|
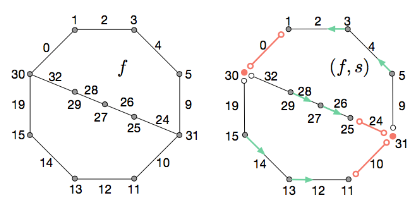
 Discrete Stratified Morse Theory: Algorithms and A User's Guide
Discrete Stratified Morse Theory: Algorithms and A User's GuideKevin Knudson and Bei Wang. Discrete & Computational Geometry, accepted, 2021. |
|
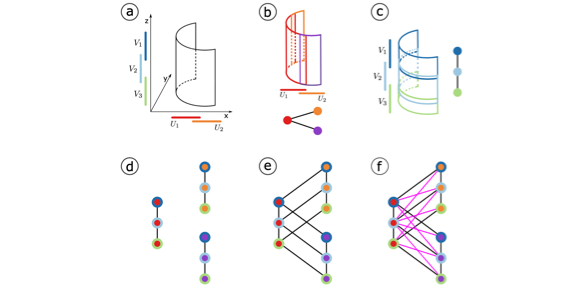
 Stitch Fix for Mapper and Topological Gains.
Stitch Fix for Mapper and Topological Gains.
Youjia Zhou, Nathaniel Saul, Ilkin Safarli, Bala Krishnamoorthy, Bei Wang. Research in Computational Topology 2, accepted, 2021. |
|
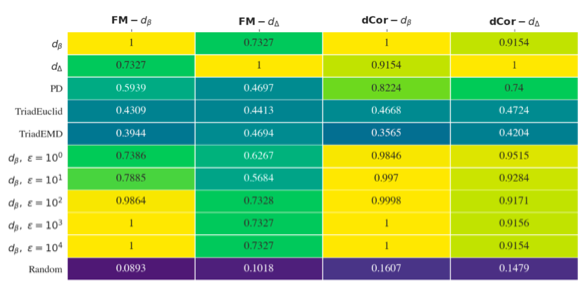
 Graph Pseudometrics from a Topological Point of View.
Graph Pseudometrics from a Topological Point of View.
Ana Lucia Garcia-Pulido, Kathryn Hess, Jane Tan, Katharine Turner, Bei Wang, Naya Yerolemou. Research in Computational Topology 2, accepted, 2021. |
|
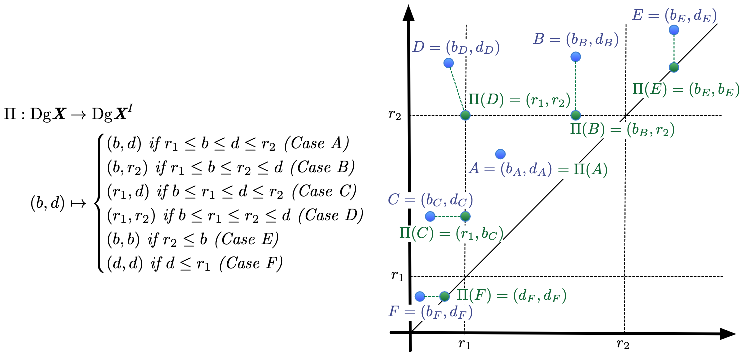
 Local Versus Global Distances for Zigzag Persistence Modules.
Local Versus Global Distances for Zigzag Persistence Modules.
Ellen Gasparovic, Maria Gommel, Emilie Purvine, Radmila Sazdanovic, Bei Wang, Yusu Wang, Lori Ziegelmeier. Research in Computational Topology 2, accepted, 2021. arXiv:1903.08298. |
|
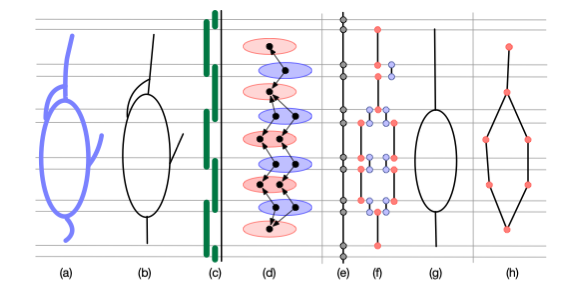
 Probabilistic Convergence and Stability of Random Mapper Graphs.
Probabilistic Convergence and Stability of Random Mapper Graphs.
Adam Brown, Omer Bobrowski, Elizabeth Munch, Bei Wang. Journal of Applied and Computational Topology, 5, pages 99-140, 2021. DOI:10.1007/s41468-020-00063-x arXiv:1909.03488. |
|
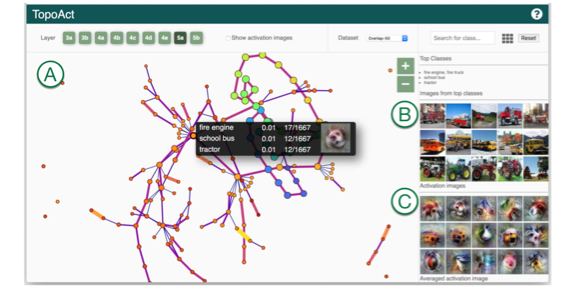
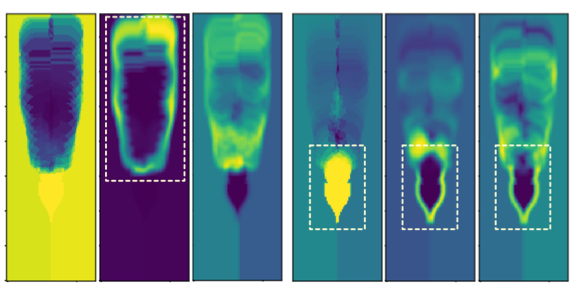
 TopoAct: Visually Exploring the Shape of Activations in Deep Learning.
TopoAct: Visually Exploring the Shape of Activations in Deep Learning.Archit Rathore, Nithin Chalapathi, Sourabh Palande, Bei Wang. Computer Graphics Forum, 40(1), pages 382-397, 2021. Supplemental Material. DOI: 10.1111/cgf.14195 arXiv:1912.06332. |
|
 Uncertainty Visualization of 2D Morse Complex Ensembles Using
Statistical Summary Maps.
Uncertainty Visualization of 2D Morse Complex Ensembles Using
Statistical Summary Maps.
Tushar Athawale, Dan Maljovec, Lin Yan, Chris R. Johnson, Valerio Pascucci, Bei Wang. IEEE Transactions on Visualization and Computer Graphics, 2020. DOI: 10.1109/TVCG.2020.3022359 |
| Year 3 (2019 - 2020) | |
|
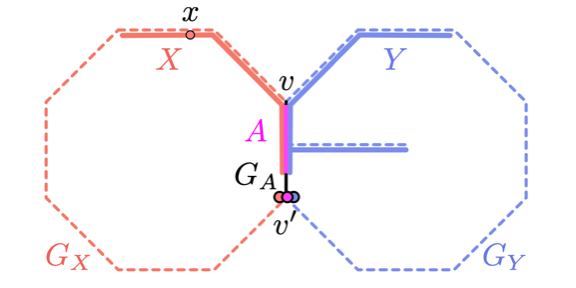
 On Homotopy Types of Vietoris--Rips Complexes of Metric Gluings.
On Homotopy Types of Vietoris--Rips Complexes of Metric Gluings.
Michal Adamaszek, Henry Adams, Ellen Gasparovic, Maria Gommel, Emilie Purvine, Radmila Sazdanovic, Bei Wang, Yusu Wang and Lori Ziegelmeier. Journal of Applied and Computational Topology, 4, pages 425-454, 2020. DOI:10.1007/s41468-020-00054-y arXiv:1712.06224. |
|
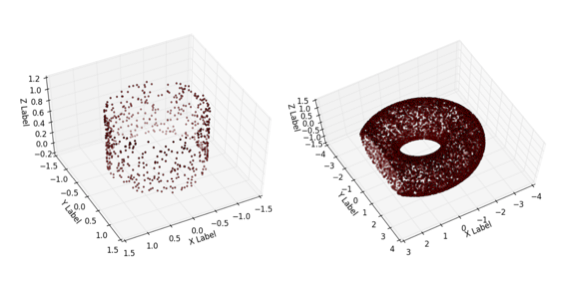
 Topological Inference of Manifolds with Boundary.
Topological Inference of Manifolds with Boundary.
Yuan Wang, Bei Wang. Computational Geometry: Theory and Applications, 88(101606), 2020. DOI:10.1016/j.comgeo.2019.101606 arXiv:1810.05759 |
|
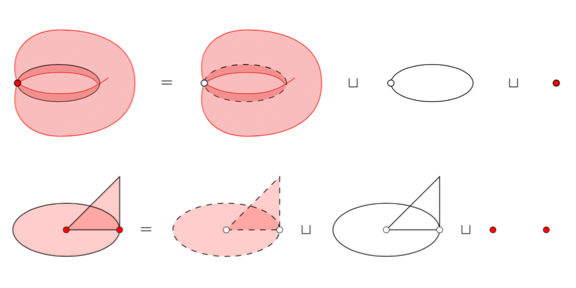
 Sheaf-Theoretic Stratification Learning From Geometric and Topological Perspectives.
Sheaf-Theoretic Stratification Learning From Geometric and Topological Perspectives.
Adam Brown and Bei Wang. Discrete & Computational Geometry, 2020. DOI:10.1007/s00454-020-00206-y arXiv:1712.07734 |
|
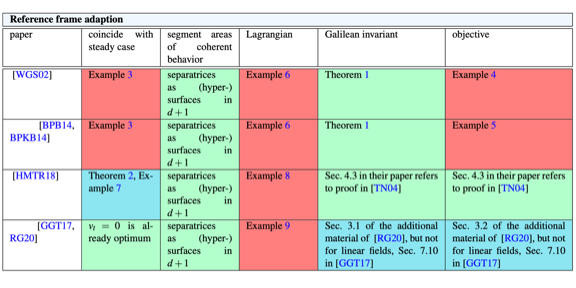
 State of the Art in Time-Dependent Flow Topology: Interpreting Physical Meaningfulness Through Mathematical Properties.
State of the Art in Time-Dependent Flow Topology: Interpreting Physical Meaningfulness Through Mathematical Properties.
Roxana Bujack, Lin Yan, Ingrid Hotz, Christoph Garth, Bei Wang. Eurographics Conference on Visualization (EuroVis) STAR Computer Graphics Forum, 2020. DOI: 10.1111/cgf.14037 |
|
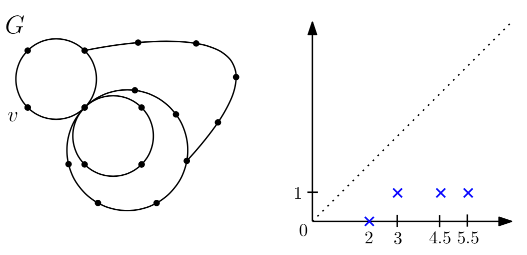
 The Relationship Between the Intrinsic Cech and Persistence Distortion Distances for Metric Graphs.
The Relationship Between the Intrinsic Cech and Persistence Distortion Distances for Metric Graphs.
Ellen Gasparovic, Maria Gommel, Emilie Purvine, Radmila Sazdanovic, Bei Wang, Yusu Wang, Lori Ziegelmeier. Journal of Computational Geometry, 10(1), pages 477-499, 2019. DOI: 10.20382/jocg.v10i1a16 arXiv:1812.05282 |
| Year 2 (2018 - 2019) | |
|
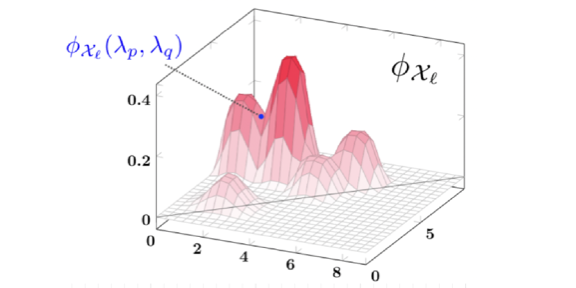
 A Kernel for Multi-Parameter Persistent Homology.
A Kernel for Multi-Parameter Persistent Homology.
René Corbet, Ulderico Fugacci, Michael Kerber, Claudia Landi, Bei Wang. Shape Modeling International (SMI), 2019. Computers & Graphics: X, 2019. arXiv:1809.10231. Best Paper Award at SMI 2019! |
|
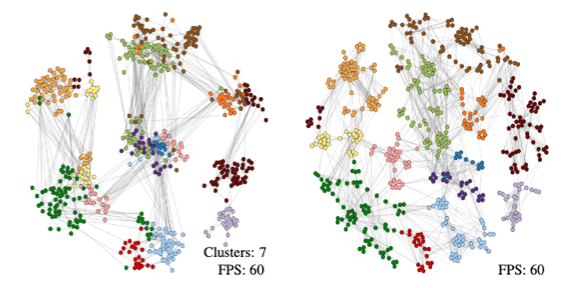
 Persistent Homology Guided Force-Directed Graph Layouts.
Persistent Homology Guided Force-Directed Graph Layouts.
Ashley Suh, Mustafa Hajij, Bei Wang, Carlos Scheidegger, Paul Rosen IEEE Transactions on Visualization and Computer Graphics (TVCG, Proceedings of InfoVis), 26(1), pages 697-707, 2020. DOI: 10.1109/TVCG.2019.2934802 arXiv:1712.05548 Long Video. Short Video. |
|
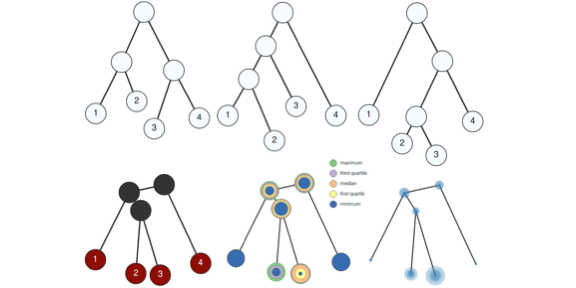
 A Structural Average of Labeled Merge Trees for Uncertainty Visualization.
A Structural Average of Labeled Merge Trees for Uncertainty Visualization.
Lin Yan, Yusu Wang, Elizabeth Munch, Ellen Gasparovic, Bei Wang. IEEE Transactions on Visualization and Computer Graphics (TVCG, Proceedings of SciVis), 26(1), pages 832-842, 2020. Supplemental Material. Doi: 10.1109/TVCG.2019.2934242 arXiv:1908.00113 Long Video. Short Video. |
| Year 1 (2017 - 2018) | |
|
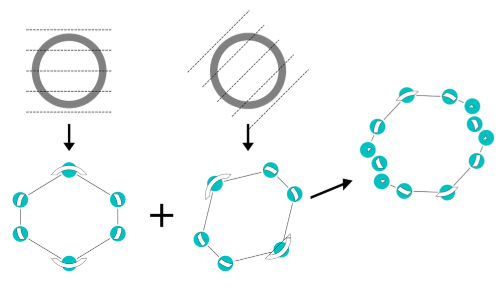
 Stitch Fix for Mapper (Abstract).
Stitch Fix for Mapper (Abstract).
Bala Krishnamoorthy, Nathaniel Saul and Bei Wang. Computational Geometry: Young Researchers Forum at International Symposium on Computational Geometry (SOCG), 2018. YRF Book of Abstracts |

|
 Discrete Stratified Morse Theory: A User's Guide.
Discrete Stratified Morse Theory: A User's Guide.
Kevin Knudson and Bei Wang. International Symposium on Computational Geometry (SOCG), 2018. DOI: 10.4230/LIPIcs.SoCG.2018.54 arXiv:1801.03183 |
|
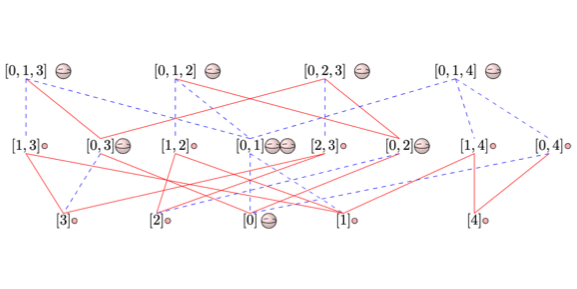
 Sheaf-Theoretic Stratification Learning.
Sheaf-Theoretic Stratification Learning.
Adam Brown and Bei Wang. International Symposium on Computational Geometry (SOCG), 2018. DOI: 10.4230/LIPIcs.SoCG.2018.14 arXiv:1712.07734 |

|
 Vietoris-Rips and Čech Complexes of Metric Gluings.
Vietoris-Rips and Čech Complexes of Metric Gluings.
Michal Adamaszek, Henry Adams, Ellen Gasparovic, Maria Gommel, Emilie Purvine, Radmila Sazdanovic, Bei Wang, Yusu Wang and Lori Ziegelmeier. International Symposium on Computational Geometry (SOCG), 2018. DOI: 10.4230/LIPIcs.SoCG.2018.3 arXiv:1712.06224 |
| Manuscripts | |

|
 Gazing Into the Metaboverse: Automated Exploration and Contextualization of Metabolic Data.
Gazing Into the Metaboverse: Automated Exploration and Contextualization of Metabolic Data.
Jordan A. Berg, Youjia Zhou, T. Cameron Waller, Yeyun Ouyang, Sara M. Nowinski, Tyler Van Ry, Ian George, James E. Cox, Bei Wang, Jared Rutter. Manuscript, 2020. bioRxiv:10.1101/2020.06.25.171850v1. |
|
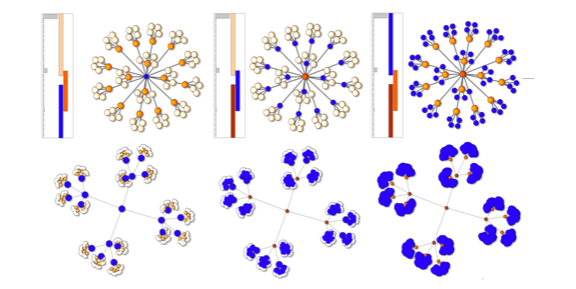
 Mapper on Graphs for Network Visualization.
Mapper on Graphs for Network Visualization.
Mustafa Hajij, Bei Wang, Paul Rosen. Manuscript, 2018. arXiv:1804.11242. |
|
|
|
Software Downloads |
|
|

|
Mapper Interactive https://mapperinteractive.github.io/ Mapper Interactive provides a library of easily-extendable modules for developing interactive visualization of high-dimensional data using the mapper construction. It is fairly lightweight, and helps to rapidly explore the parameter space of the mapper construction interactively. Mapper Interactive is under active development. Please excuse its appearance (e.g., documentations). |

|
Pheno-Mapper https://github.com/tdavislab/PhenoMapper Pheno-Mapper is a domain-specific adaptation of Mapper Interactive and specifically targets the analysis and visualization of multi-dimensional phenomics data. |
|
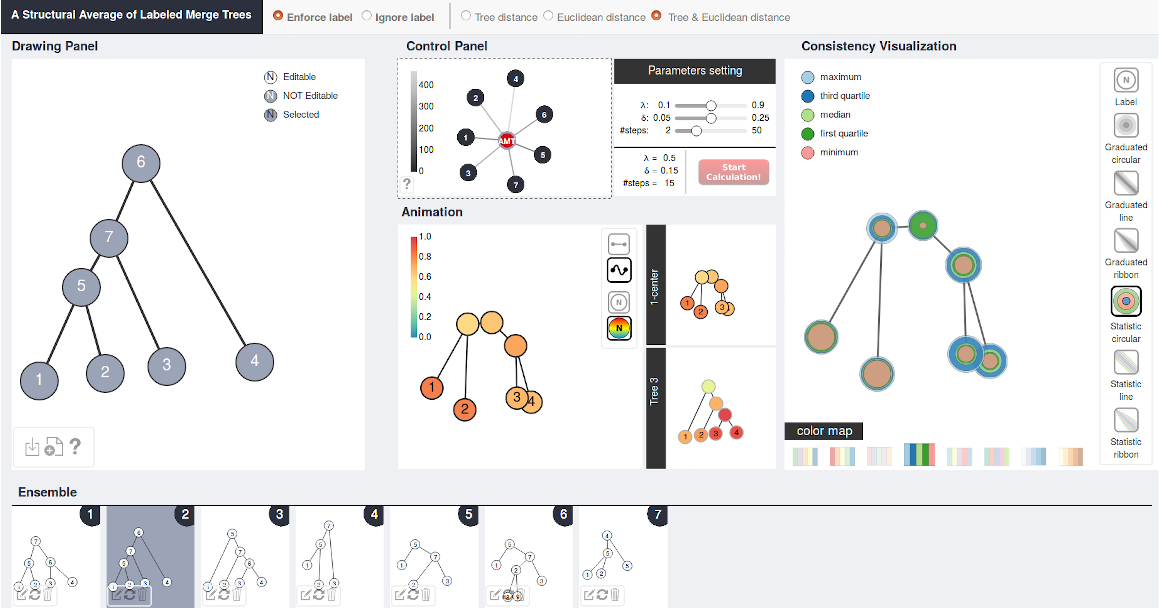
AMT: Interactive Visualization of Labeled Merge Trees and Their 1-Center https://github.com/tdavislab/amt AMT is an interactive visualization tool that computes an average tree structure for an ensemble of input (merge) trees. It can also be potentially used to compute an average clustering given a collection of hierarchical clustering results. AMT is under active development. Please excuse its appearance (e.g., documentations). |

|
Metaboverse: Automated Exploration and Contextualization of Metabolic Data.
https://github.com/Metaboverse/ Metaboverse is an interactive visualization tool for automated exploration and contextualization of metabolic data. Integrating multi-omic or single-omic metabolic data upon the metabolic network can be challenging for a variety of reasons. Metaboverse seeks to simplify this task for users by providing a simple, user-friendly interface for layering their data on a dynamic representation of the metabolic network. Additionally, it provides several new tools to enable the contextualization of metabolic data. Metaboverse is under active development. Please excuse its appearance (e.g., documentations). |

|
TopoAct: Visually Exploring the Shape of Activations in Deep Learning. https://github.com/tdavislab/TopoAct/ TopoAct is a visual exploration system used to study topological summaries of activation vectors for deep neural networks. We present visual exploration scenarios using TopoAct that provide valuable insights towards learned representations of image classifiers such as GoogLeNet and ResNet. TopoAct is under active development. |
|
|
|
Presentations, Educational Development and Broader Impacts |
|
|
| Year 4 (2020 - 2021) |
|
Nithin Chalapathi Conference Talk (virtual): Adaptive Covers for Mapper Graphs Using Information Criteria at IEEE International Conference on Big Data: Workshop on Applications of Topological Data Analysis to Big Data, December 15, 2021.
|
| Year 3 (2019 - 2020) |
|
Bei Wang Invited Talk (virtual): Topology as a knob for machine learning at MBI Optimal Transport Workshop: Optimal Transport, Topological Data Analysis and Applications to Shape and Machine Learning, July 27-31, 2020. |
| Year 2 (2018 - 2019) |
|
Bei Wang and Bala Krishnamoorthy Workshop Organization:
8th Annual Minisymposium on Computational Topology during the Computational Geometry Week, June 18-21, 2019. |
| Year 1 (2017 - 2018) |
|
Bei Wang Conference Talk: Discrete Stratified Morse Theory: A User's Guide, at 34th International Symposium on Computational Geometry (SOCG), June 11-14, 2018. |
|
|
|
Students |
|
|
|
Lin Yan (Fall 2017 - Present) School of Computing and Scientific Computing and Imaging Institute University of Utah linyan AT sci.utah.edu Youjia Zhou (Spring 2019 - Present) School of Computing and Scientific Computing and Imaging Institute University of Utah zhou325 AT sci.utah.edu Fangfei Lan (Spring 2020 - Present) School of Computing and Scientific Computing and Imaging Institute University of Utah fangfei.lan AT sci.utah.edu Archit Rathore (Summer 2018 - Present) School of Computing and Scientific Computing and Imaging Institute University of Utah archit.rathore AT utah.edu Ilkin Safarli (Summer 2020 - Spring 2021, graduated Summer 2021) School of Computing and Scientific Computing and Imaging Institute University of Utah Yaodong Zhao (Fall 2017 - Spring 2019, graduated Spring 2019) School of Computing and Scientific Computing and Imaging Institute University of Utah |
|
|
|
Acknowledgement |
|
|
|
This material is based upon work supported or partially supported by the National Science Foundation under Grant No. 1661375 and 1661348, project titled "ABI Innovation: A Scalable Framework for Visual Exploration and Hypotheses Extraction of Phenomics Data using Topological Analytics." |
|
|