2015
I. OguzI, J. Cates, M. Datar, B. Paniagua, T. Fletcher, C. Vachet, M. Styner, R. Whitaker.
“Entropy-based particle correspondence for shape populations,” In International Journal of Computer Assisted Radiology and Surgery, Springer, pp. 1-12. December, 2015.
Purpose
Statistical shape analysis of anatomical structures plays an important role in many medical image analysis applications such as understanding the structural changes in anatomy in various stages of growth or disease. Establishing accurate correspondence across object populations is essential for such statistical shape analysis studies.
Methods
In this paper, we present an entropy-based correspondence framework for computing point-based correspondence among populations of surfaces in a groupwise manner. This robust framework is parameterization-free and computationally efficient. We review the core principles of this method as well as various extensions to deal effectively with surfaces of complex geometry and application-driven correspondence metrics.
Results
We apply our method to synthetic and biological datasets to illustrate the concepts proposed and compare the performance of our framework to existing techniques.
Conclusions
Through the numerous extensions and variations presented here, we create a very flexible framework that can effectively handle objects of various topologies, multi-object complexes, open surfaces, and objects of complex geometry such as high-curvature regions or extremely thin features.
2013
M. Datar, I. Lyu, S. Kim, J. Cates, M.A. Styner, R.T. Whitaker.
“Geodesic distances to landmarks for dense correspondence on ensembles of complex shapes,” In Proceedings of Medical Image Computing and Computer-Assisted Intervention (MICCAI 2011), Vol. 16(Pt. 2), pp. 19--26. 2013.
PubMed ID: 24579119
Establishing correspondence points across a set of biomedical shapes is an important technology for a variety of applications that rely on statistical analysis of individual subjects and populations. The inherent complexity (e.g. cortical surface shapes) and variability (e.g. cardiac chambers) evident in many biomedical shapes introduce significant challenges in finding a useful set of dense correspondences. Application specific strategies, such as registration of simplified (e.g. inflated or smoothed) surfaces or relying on manually placed landmarks, provide some improvement but suffer from limitations including increased computational complexity and ambiguity in landmark placement. This paper proposes a method for dense point correspondence on shape ensembles using geodesic distances to a priori landmarks as features. A novel set of numerical techniques for fast computation of geodesic distances to point sets is used to extract these features. The proposed method minimizes the ensemble entropy based on these features, resulting in isometry invariant correspondences in a very general, flexible framework.
2012
B. Paniagua, L. Bompard, J. Cates, R.T. Whitaker, M. Datar, C. Vachet, M. Styner.
“Combined SPHARM-PDM and entropy-based particle systems shape analysis framework,” In Medical Imaging 2012: Biomedical Applications in Molecular, Structural, and Functional Imaging, SPIE Intl Soc Optical Eng, March, 2012.
DOI: 10.1117/12.911228
PubMed ID: 24027625
PubMed Central ID: PMC3766973
2011
M. Datar, Y. Gur, B. Paniagua, M. Styner, R.T. Whitaker.
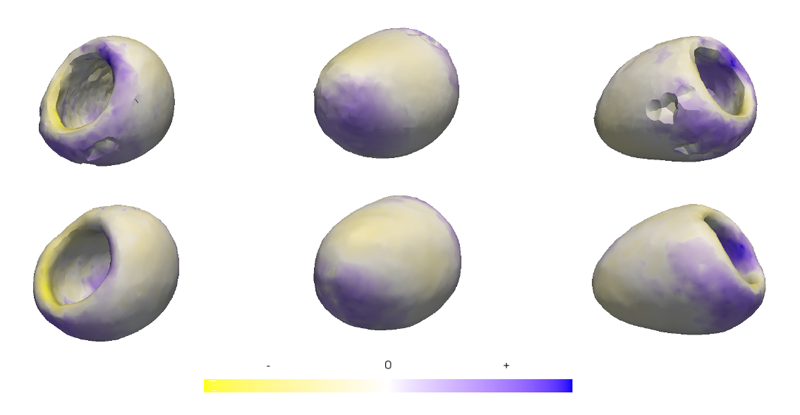
“Geometric Correspondence for Ensembles of Nonregular Shapes,” In Proceedings of Medical Image Computing and Computer-Assisted Intervention (MICCAI 2011), Lecture Notes in Computer Science (LNCS), Vol. 6892, pp. 368--375. 2011.
DOI: 10.1007/978-3-642-23629-7_45
PubMed ID: 21995050
PubMed Central ID: PMC3346950

Keywords: namic
2009
I. Oguz, M. Niethammer, J. Cates, R.T. Whitaker, P.T. Fletcher, C. Vachet, M. Styner.
“Cortical Correspondence with Probabilistic Fiber Connectivity,” In Information Processing in Medical Imaging (IPMI), Lecture Notes in Computer Science (LCNS), Vol. 5636, pp. 651--663. 2009.
DOI: 10.1007/978-3-642-02498-6_54
2008
J. Cates, P.T. Fletcher, M. Styner, H. Hazlett, R.T. Whitaker.
“Particle-Based Shape Analysis of Multi-Object Complexes,” In Proceedings of the 11th International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI '08), Lecture Notes In Computer Science (LCNS), pp. 477--485. 2008.
ISBN: 978-3-540-85987-1
I. Oguz, J. Cates, P.T. Fletcher, R.T. Whitaker, D. Cool, S. Aylward, M. Styner.
“Cortical Correspondence using Entropy-Based Particle Systems and Local Features,” In 5th IEEE International Symposium on Biomedical Imaging: From Nano to Macro, 2008. (ISBI 2008), pp. 1637--1640. 2008.
S. Pizer, M. Styner, T. Terriberry, R. Broadhurst, S. Joshi, E. Chaney, P.T. Fletcher.
“Statistical Applications with Deformable M-Reps,” In Computational Imaging and Vision, Springer, pp. 269--308. 2008.
DOI: 10.1007/978-1-4020-8658-8_9
2007
J. Cates, P.T. Fletcher, M. Styner, M. Shenton, R.T. Whitaker.
“Shape Modeling and Analysis with Entropy-Based Particle Systems,” In Proceedings of Information Processing in Medical Imaging (IPMI) 2007, LNCS 4584, pp. 333--345. 2007.