Technical Research and Development
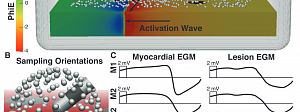
Simulation and Estimation
The simulation of bioelectric fields, and estimation of their parameters from measurements, have always been a core research domain for CIBC, and we continue to pursue exciting new topics in our Simulation & Estimation TR&D. We also continue to create and… Read more
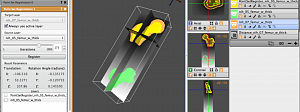
Image and Geometric Analysis
The Image and Geometric Analysis TRD addresses an ongoing demand for software that allows biomedical scientists to quickly build geometric and statistical representations from collections of images. Motivated by the needs of collaborators and the technical… Read more
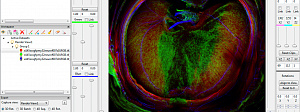
Visualization
Because the development of infrastructure required for conducting and analyzing volume data-intensive experiments and simulations has not kept pace with our collective ability to gather and create large-scale data, current data analysis and volume… Read more