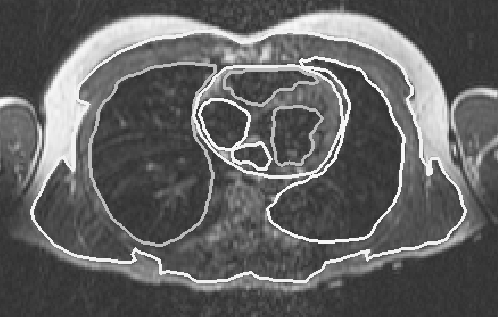
Figure 1: Magnetic resonance image of the thorax with superposed segmentation boundaries. The projection of the image is caudal-cranial and tissue boundaries include subcutaneous fat, lungs, and epicardial and endocardial borders of the heart.
The model in this study was constructed primarily from 115 transverse MRIs of a healthy adult male. The images were taken every 5 mm beginning just below the waist and ending at the bottom of the neck. In each two-dimensional image, the regions of interest were segmented into contours. Figure 1 shows the segmentation contours of several major regions in one of the images that included the heart. The segmentation was accomplished with a segmentation program that allowed the user to view the MRI to manually select boundary points with a mouse [22, 23]. The points for each contour were used to create a spline that was then resampled to establish regular spacing and saved as a set of x,y coordinates describing the contour at a particular z level. In this way, regions of particular tissue types (for example, subcutaneous fat) as well as anatomical structures (such as the right atrium) were specified. The larger tissue regions of the model included the subcutaneous fat, the skeletal muscle, the lungs and the bones of the torso. In the vicinity of the heart, more detailed anatomical structures were specified. These structures included both atria and ventricles, three epicardial fat pads and the following components of the great vessels: the superior and inferior vena cava, the pulmonary arteries and veins, the ascending and descending aorta, the azygous vein, and the left and right subclavian veins. All of the contours in the different z planes for a particular structure, such as a lung, were then connected into triangles to construct a closed surface.

Figure 1: Magnetic resonance image of the thorax with superposed segmentation boundaries. The projection of the image is caudal-cranial and tissue boundaries include subcutaneous fat, lungs, and epicardial and endocardial borders of the heart.
For each closed surface, a Delaunay optimal tetrahedral mesh was then generated to discretize the volume of the thorax (the mesh generator used for this study is described in detail elsewhere [24, 25]). Tetrahedra were chosen because of their ability to approximate the irregular surfaces and volumes of the human body and because they are well suited to fully automatic mesh generation algorithms. As part of the tetrahedralization procedure, each element also received a group number based on the tissue type assigned to the cell. This group number later served as the means of flexibly assigning conductivity values to each element in the model. Figure 2 shows a cutaway of the outer torso surface with the triangulated surfaces of the lungs and the epicardium inside.

Figure 2: Frontal view of the surface triangulation of the Utah torso model. Surfaces included in the figure are torso boundary, lungs, and epicardium. Internodal spacing ranged from 5-10 mm.
In a second iteration of constructing the Utah Torso, we incorporated the following bones into the torso model: the left and right clavicle, the sternum, all 12 left and right ribs, and the spinal column. Because the bones were not well defined in the MRIs, surface geometry for these structures was obtained from Viewpoint (Orem, Utah), a commercial supplier of three-dimensional geometry data sets. The point coordinates were scaled and shifted to fit within the Utah Torso. The mesh of the bones was then generated with the same code used for the structures obtained from the MRIs. The complete torso mesh had approximately one million tetrahedra and 168,000 nodes. The simulations in this study were all calculated using a single mesh. If different meshes were used based on the different inhomogeneities being modeled in a particular scenario, the effects calculated would be even greater.