 |
Rob MacLeod
The programs described in this document all serve one purpose--to make management of needle geometry as easy as possible. They take care of all the steps required to convert digitized rod entry points and rod tips into geometry files, with a few additional ``goodies'' for fun.
The steps required to manage needles are as follows:
The programs are not perfect, nor is the documentation, but we can make it that way with your help. Let Rob know what is not clear or what fails to perform as required and we will aim to please!
The format of data that comes from the digitizer is not directly usable by any of the standard CVRTI programs and must be converted before anything else happens with is. The program digtoxyz is one program for this conversion. It takes digitized datasets in a variety of formats and tries to identify what the contents are, then converts and saves them in the appropriate manner. The reason for the complexity lies in the diversity of things we digitize, from sock electrodes points to the tips and rods, and the need to treat each one a little differently. A further complexity lies in the two different devices we use to digitize; on good days this program handles both with grace and beauty.
Below we list each of the different types of geometrical objects or information that digtoxyz knows about and what type of input file format is necessary for consistent interpretation. I have removed all references to what used to be called the ``old digitizer'', this was the one built at the CVRTI that fixes the object on a rotating jig and moves the caliper to touch the points. We no longer use nor support this device. In contrast, the ``Microscribe digitizer'' is the device we have used for 10 years now. We create files from this device using Phil's program on the Macintosh. The standard extension for these files is ``.ms''.
There is no magic nor strict paradigm for digitizing with the Microscribe; on the other hand, a few lessons from experience can be useful:
The program digtoxyz does not explicitly care about the order of items digitized, nor which itens are included in a single .ms file. However, a little planning in this regard can save time in the subsequenct processing.
Crucial in the entry of needle/rod points is the order in which the points of each needle/rod get digitized. The convention as we have agreed to it so far is outer end = rod tip, then inner end = rod entry point.
To run the program digtoxyz requires the following command line:
digtoxyz [-m -c] infilename outfilename
where
Note: New digitizer includes the units in the file while the old digitizer defaults to inches for units.
Sock electrode locations are read as a series of points, and then later triangulated and formed into geometric meshes that form the basis for visualization, computer models and other types of data analysis. Hence the role of digtoxyz is fairly simple--convert each point from digitizer format to a geometry file (either .geom or .pts/.fac files).
| Header Lines (4 in all) | |||
| FILE NAME: filename LAST MODIFIED: Date and Time | |||
| Free line of text | |||
| ``Measurements are in'' millimeters or inches) | |||
| **************************************** | |||
| File Body | |||
| ``Point'' # | X-value | Y-value | Z-value |
| Example | |||
| zero | 0.000 | 0.000 | 0.000 |
| sock 1 | 28.508 | -173.895 | 221.950 |
| sock 2 | 20.061 | -178.593 | 225.062 |
| sock 3 | 11.862 | -179.161 | 237.611 |
| sock 4 | 5.729 | -175.170 | 251.998 |
The result of running digtoxyz on epicardial points is a file that is part of the CVRTI geometry files. Its extensions is .pts and it contains a list of all the points, converted into Cartesian coordinates and units of millimeters.
These are the files that result from digitizing the paths of the coronary arteries, which needs to be done according to the following tips:
There are two slightly different file layouts that digtoxyz will recognize as being from a coronary artery tree. One contains extra information of a radius for each point in the tree. This eventually determines the thickness of the tubes drawn in map3d and if fixed at the time of digitization, then it supplies one set of values that digtoxyz otherwise requires from the user when it runs.
The first format includes only the coordinates of each artery point.
| Header Lines (4 in all) | |||
| FILE NAME: filename LAST MODIFIED: Date and Time | |||
| Free line of text | |||
| ``Measurements are in'' millimeters or inches) | |||
| **************************************** | |||
| File Body | |||
| ``Point'' # | X-value | Y-value | Z-value |
| Example | |||
| Artery 1-1 | 28.508 | -173.895 | 221.950 |
| Artery 1-2 | -.9385 | 4.8480 | 225.062 |
| Artery 1-3 | -1.0225 | 4.5970 | 237.611 |
| Artery 1-4 | -1.1590 | 4.4565 | 251.998 |
| Artery 1-5 | -1.2280 | 4.2110 | 221.950 |
| Artery 1-6 | -1.2645 | 3.9330 | 225.062 |
| Artery 1-7 | -1.2510 | 3.6935 | 237.611 |
| Artery 1-8 | -1.2050 | 3.5000 | 251.998 |
| Artery 2-1 | -.9345 | 4.8945 | 221.950 |
| Artery 2-2 | -.8620 | 4.5170 | 225.062 |
| Artery 2-3 | -.7850 | 4.2745 | 237.611 |
| Artery 2-4 | -.6610 | 3.9005 | 251.998 |
| Artery 2-5 | -.5445 | 3.6085 | 221.950 |
The second format includes a radius value for each point, something one must typically add manually.
| Header Lines (4 in all) | ||||
| FILE NAME: filename LAST MODIFIED: Date and Time | ||||
| Free line of text | ||||
| ``Measurements are in'' millimeters or inches) | ||||
| **************************************** | ||||
| File Body | ||||
| ``Point'' # | X-value | Y-value | Z-value | Radius |
| Example | ||||
| Artery 1-1 | 28.508 | -173.895 | 221.950 | .25 |
| Artery 1-2 | -.9385 | 4.8480 | 225.062 | .15 |
| Artery 1-3 | -1.0225 | 4.5970 | 237.611 | .14 |
| Artery 1-4 | -1.1590 | 4.4565 | 251.998 | .14 |
| Artery 1-5 | -1.2280 | 4.2110 | 221.950 | .13 |
| Artery 1-6 | -1.2645 | 3.9330 | 225.062 | .1 |
| Artery 1-7 | -1.2510 | 3.6935 | 237.611 | .05 |
| Artery 1-8 | -1.2050 | 3.5000 | 251.998 | .01 |
| Artery 2-1 | -.9345 | 4.8945 | 221.950 | .15 |
| Artery 2-2 | -.8620 | 4.5170 | 225.062 | .12 |
| Artery 2-3 | -.7850 | 4.2745 | 237.611 | .1 |
| Artery 2-4 | -.6610 | 3.9005 | 251.998 | .05 |
| Artery 2-5 | -.5445 | 3.6085 | 221.950 | .01 |
The output file for coronaries has to be more complicated than the simple list of points that results from the epicardial sock points. The file format used for this information is called a landmark file and the details of its purpose and format are in the manual for map3d. Briefly, this is another simple text file organize by landmark type and segments. There are several different landmark types, one of which is a coronary segment, that consists of a list of all the digitized points, ordered according to the numbers supplied in the digitizer file.
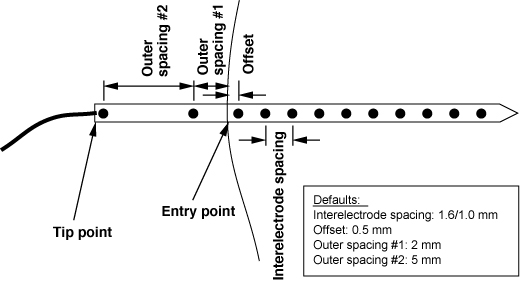
Needles are usually digitized by replacing each needle with a rod of known length and then digitizing the tips and entry points of each needle. From this information and knowledge of the needle electrode size and number of electrodes we can construct the locations of all the needle electrodes. Recently, we have used two points on the needle to determine direction, removing the need for rods to replace each needle. For historic and compatibilty reasons, however, the key word in the .ms file for both rods and needles is ``rod''. The order assumed for the points on each rod or needle is outer end = rod tip, then inner end = rod entry point.
The first step is to convert the entry points and tips into a landmarks file that becomes the input to both map3d and subsequent processing programs described in Section 3 of this document..
| Header Lines (4 in all) | |||
| FILE NAME: filename LAST MODIFIED: Date and Time | |||
| Free line of text | |||
| Measurements are in millimeters (or inches) | |||
| **************************************** | |||
| File Body | |||
| ``Rod'' #-1/2 | X-value | Y-value | Z-value |
| Example | |||
| Rod 1-1 | 28.508 | -173.895 | 221.950 |
| rod 1-2 | 20.061 | -178.593 | 225.062 |
| rod 2-1 | 11.862 | -179.161 | 237.611 |
| Rod 2-2 | 5.729 | -175.170 | 251.998 |
Notes:
The output file from digtoxyz is a landmarks file, this time with objects of type rod. This file is already in the form that map3d uses so that quality control the digitization and conversion is quite easy. For example, with a points file from the epicardial surface called 24mar96_sock.pts and a rod landmarks file called 24mar96_rods.lmarks a sample command line for map3d would be:
map3d -f 24mar96_sock -lm 24mar96-rods.lmarks
The main utility of the rods landmarks file, however, is to convert it into needle electrode positions, for which we have the program needlegeom , described in section 3 of this document.
Most single locations are organized in a similar fashion in digtoxyz , as is the resulting landmarks file, which is the output format for such sites.
| Header Lines (4 in all) | |||
| FILE NAME: filename LAST MODIFIED: Date and Time | |||
| Free line of text | |||
| Measurements are in millimeters (or inches) | |||
| **************************************** | |||
| File Body | |||
| Point-type # | X-value | Y-value | Z-value |
| Example | |||
| occl 1 | 28.508 | -173.895 | 221.950 |
| occl 2 | 20.061 | -178.593 | 225.062 |
| pace 1 | 11.862 | -179.161 | 237.611 |
| infusion 1 | 5.729 | -175.170 | 251.998 |
| lead 1 | 9.729 | -134.170 | 123.998 |
Note the example contains names for each of the various single points that we currently recognize. ``occl'' = occlusion; ``pace'' = pacing site, ``infus'' = infusion site; and ``lead'' = selected lead. These items can appear as part of other digitizer files, eg., at the end of a file containing coronaries.
The output file is a landmarks file. A single landmarks file can contain any mix of the different objects we have described so far. Hence you may wish to digitize coronaries and single points on one file, then have a separate file for needles, and a third file for sock electrodes.
The purpose of needlegeom is to convert the contents of a landmarks file containing rod or needle endpoints into a .geom file containing needle electrode locations. This output file then serves as input for map3d and any program that reads needle geometry.
To perform this task, needlegeom needs some information, some of which is part of the command line, with the rest (optionall) coming from supplementary files. The command line arguments are as follows:
needlegeom lmarkfile geomfile
[-i needleinfofile]
[-n num_elecs_per_needle]
[-e elec_spacing(mm)]
[-o offset] [-c channeloffset]
[-t transmatrixfilename]
If you have a needle info file for the experiment dated 24mar96, the command would be :
needlegeom 24mar96-needles.lmarks 24mar96-needles.geom -i 24mar96.ni
We assume that the landmarks file 24mar96-needles.lmarks already
exists and that the output file 24mar96-needles.geom does not (or
you wish to overwrite it).
The file 24mar96.ni is the needle
info file, which we describe next.
If we had a simple case in which all needles in the landmarks file followed in order, and there were 10 electrodes per needle, with otherwise default values the command line would become
needlegeom 24mar96-needles.lmarks 24mar96-needles.geom -e 10or if needles began at channel 193
needlegeom 24mar96-needles.lmarks 24mar96-needles.geom -e 10 -o 192Note that the value after the -o option is an offset applied to all channel numbers so for offset = 192, the first needle channel is 193.
A needle info file serves a number of purposes in telling needlegeom :
 |
The format of the needleinfo file is an ASCII, or plain text file, organized as follows:
| Format of needle info file | |
| Comment lines: | |
| # A comment | |
| Header Line: | |
| numneedles | Number of needles in the file |
| Subsequent lines | |
| First field | Needle label number |
| Second field | Channel number of distal tip electrode |
| Third field | Number of electrodes per needle |
| Fourth field | Spacing distances, in mm: interelectrode [outer_1 outer_2] |
| Fifth field | Offset distance in mm to first electrode |
| Sample Needle Info File | |||||
| # A sample needle info file for you | |||||
| # It contains lots of variety | |||||
| 14 | 14 needles described in the file | ||||
| 1 | 321 | 10 | 1.2 | 0.5 | First needle begins at channel 321, so runs from 321-330 |
| 2 | 331 | 10 | 1.2 | .5 | Second needle begins at channel 331, so runs from 331-340 |
| 3 | 341 | 10 | 1.2 | 0.5 | There are 10 electrode per needle and 1.2 mm spacing |
| 4 | 351 | 3 | 1.6 | 0.5 | Only three electrodes in this needle |
| 5 | 354 | 10 | 1.2 | 0.5 | Note the jump in channel # of only three from the last needle |
| 6 | 364 | 10 | 1.2 | 0.5 | |
| 8 | 384 | 10 | 1.2 | 0.5 | Jump in needle label; # 7 fell out |
| 9 | 394 | 10 | 1.2 | 0.5 | |
| 10 | 404 | 12 | 1.6 2.0 5.0 | 0.5 | 12-pole needle, spacing changes to 1.6 2.0 amd 5.0 mm |
| 11 | 416 | 12 | 1.6 2.0 5.0 | ||
| 12 | 428 | 12 | 1.6 2.0 5.0 | 0.5 | |
| 13 | 438 | 12 | 1.6 2.0 5.0 | 0.5 | Channels jumps by only 10; last 2 electrodes of previous needle not wired |
| 15 | 448 | 10 | 1.6 | 0.5 | Skipped another needle label |
| 16 | 458 | 10 | 1.6 | 0.5 | 14 needles but last one has label 16 |
To create these files, use any editor (if you use Microsoft Word, be sure to save the file as text only, without line-breaks) and then transfer the file to the SGI directory where the landmarks and geometry files live. The standard extension for the needleinfo file is .ni or .needleinfo. 1
The program to place epicardial sock geometry into the reference frame of the torso tank is called align. More generally, align will align any two sets of points according to some subset of points from both files that should be approximately the same. It generates a transformation matrix that any program can multiply with the nodes of the geometry to apply the same transformation to any other points. When we run align we apply the matrix automatically to the nodes of the geometry, and any landmarks in a file optionally included on the command line. Usually, we don't include the rod points in this landmarks file (although this would work too) yet we need to apply the same transformation to the needle points that we apply to the sock (and coronaries, stimulus sites, or any other landmarks that we digitize at the same time as the sock). Thus, the transformation matrix.
The format of the transformation is very simple, and the align program actually generates the text for this file, and writes it at the time of execution to a log file for your convenience. Just make sure the file exists, and the include it in the argument list for needlegeom .
The program needletri has a simple purpose--to take the triangles already entered for surfaces of a needle geometry and propagate them through a subset of the other surfaces in that geometry.
It is effectively the final stage in going from digitized rods to a needle geometry and other sections of this document describe the other steps.
The command line for needletri is as follows:
needletri infilename outfilename
where infilename is the name of the input geometry file and outfilename the name of the desired output filename (which can be the same
as the input).
All that is necessary is to have at least one surface in the input file triangulated and the program applies this same triangularization to any other surface in the file. The user controls which surfaces get propagated to which others and the program does some sanity checks for the number of points in each surface (i.e., you cannot take a triangulated surface with 10 points and propagate it to a surface with 12 points). Another assumption is that we have a regular array of needles with little or no crossing of needles within the tissue. But in the worst case, some manual editing of the resulting file will be necessary with map3d.
The purpose of the makeplanes program is to extract sets of needles from a multi-needle geometry file and put them together in a separate file for display of data.
makeplanes in.geom out.geom config.ni [-o] [-c]
where
in.geom is the input geometry file
out.geom is the desired output geometry file
config.ni is the needle info file
-o = assign channels in .ni file to outermost electrode
-c = close the ends of the planes
In more detail
Note that the input file will have multiple surfaces, one for each of the electrodes in the needles. The output file, on contrast, has just one surface and the triangulation of this file is across the needles, in a sense orthogonal to the triangulation of the input file.
makeplanes bt11may94.geom two24-1092.geom two24-1092.ni -oThe famous case which initiated the -o option. This one takes needles from the file bt11may94.geom according to the contents of two24-1092.ni and makes a new file called two24-1092.geom. The -o option ensures that the electrode order along the needle is reversed, i.e., that the channel numbers in two24-1092.ni refer to the outermost electrode.
The file two24-1092.ni looks like this
9 1 24 12 1.6 2.0 4.0 0.5 2 192 12 1.6 2.0 4.0 0.5 3 336 12 1.6 2.0 4.0 0.5 4 408 12 1.6 2.0 4.0 0.5 5 600 12 1.6 2.0 4.0 0.5 6 504 12 1.6 2.0 4.0 0.5 7 888 12 1.6 2.0 4.0 0.5 8 1032 12 1.6 2.0 4.0 0.5 9 1092 12 1.6 2.0 4.0 0.5
So 24 refers to channel 24, which is on the outer end of the needle (13 is on the inner end).
This document was generated using the LaTeX2HTML translator Version 2002-2-1 (1.71)
Copyright © 1993, 1994, 1995, 1996,
Nikos Drakos,
Computer Based Learning Unit, University of Leeds.
Copyright © 1997, 1998, 1999,
Ross Moore,
Mathematics Department, Macquarie University, Sydney.
The command line arguments were:
latex2html -split 3 -no_white -link 3 -no_navigation -no_math -html_version 3.2,math -show_section_numbers -local_icons needles
The translation was initiated by Rob Macleod on 2007-06-23