News
Utah-Led Group Gets $15M from Army to Design New Materials
 May 7, 2012 – U.S. soldiers are increasingly weighed down by batteries to power weapons, detection devices and communications equipment. So the Army Research Laboratory has awarded a University of Utah-led consortium almost $15 million to use computer simulations to help design materials for lighter-weight, energy efficient devices and batteries.
May 7, 2012 – U.S. soldiers are increasingly weighed down by batteries to power weapons, detection devices and communications equipment. So the Army Research Laboratory has awarded a University of Utah-led consortium almost $15 million to use computer simulations to help design materials for lighter-weight, energy efficient devices and batteries."We want to help the Army make advances in fundamental research that will lead to better materials to help our soldiers in the field," says computing Professor Martin Berzins, principal investigator among five University of Utah faculty members who will work on the project. "One of Utah’s main contributions will be the batteries."
Of the five-year Army grant of $14,898,000, the University of Utah will retain $4.2 million for research plus additional administrative costs. The remainder will go to members of the consortium led by the University of Utah, including Boston University, Rensselaer Polytechnic Institute, Pennsylvania State University, Harvard University, Brown University, the University of California, Davis, and the Polytechnic University of Turin, Italy.
Epinome: Visual-Analytics for Epidemiology Data
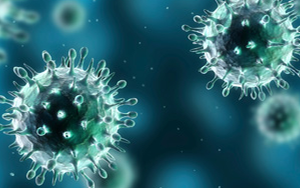 Early detection and rapid response to infectious-disease outbreaks rely on effective decision making based on information from disparate sources. To improve decision-making in outbreak detection and response, it's important to understand how public health practitioners seek relevant information. Epinome, a user-centric visual-analytics system, supports research on decision-making in public health, particularly evaluation of information search strategies. Epinome facilitates investigation of scripted high-fidelity large-scale simulated disease outbreaks. Its dynamic environment seamlessly evolves and adapts as the user's tasks and focus change. This video shows how the Epinome system facilitates interactive simulations of disease outbreaks.
Early detection and rapid response to infectious-disease outbreaks rely on effective decision making based on information from disparate sources. To improve decision-making in outbreak detection and response, it's important to understand how public health practitioners seek relevant information. Epinome, a user-centric visual-analytics system, supports research on decision-making in public health, particularly evaluation of information search strategies. Epinome facilitates investigation of scripted high-fidelity large-scale simulated disease outbreaks. Its dynamic environment seamlessly evolves and adapts as the user's tasks and focus change. This video shows how the Epinome system facilitates interactive simulations of disease outbreaks.Read Full Article (pdf)
Y. Livnat, T.-M. Rhyne, M. Samore. "Epinome: A Visual-Analytics Workbench for Epidemiology Data," In IEEE Computer Graphics and Applications, Vol. 32, No. 2, pp. 89--95. 2012. ISSN: 0272-1716 DOI: 10.1109/MCG.2012.31
Visualizing the Future of Medicine
 CIBC Software freatures prominently in Scientific Computing World's interview of Chris Johnson "Visualizing the Future of Medicine."
CIBC Software freatures prominently in Scientific Computing World's interview of Chris Johnson "Visualizing the Future of Medicine." See: Scientific Computing World - "Visualizing the Future of Medicine" (pdf version)
U.S. Energy Secretary Steven Chu Announces SDAV
 Energy Secretary Steven Chu announced a $25 million five-year initiative to help scientists better extract insights from today’s increasingly massive research datasets, the Scalable Data Management, Analysis, and Visualization (SDAV) Institute. SDAV will be funded through DOE’s Scientific Discovery through Advanced Computing (SciDAC) program and led by Arie Shoshani of Lawrence Berkeley National Laboratory (Berkeley Lab). As one of the nation’s leading funders of basic scientific research, the Department of Energy Office of Science has a vested interested in helping researchers effectively manage and use these large datasets.
Energy Secretary Steven Chu announced a $25 million five-year initiative to help scientists better extract insights from today’s increasingly massive research datasets, the Scalable Data Management, Analysis, and Visualization (SDAV) Institute. SDAV will be funded through DOE’s Scientific Discovery through Advanced Computing (SciDAC) program and led by Arie Shoshani of Lawrence Berkeley National Laboratory (Berkeley Lab). As one of the nation’s leading funders of basic scientific research, the Department of Energy Office of Science has a vested interested in helping researchers effectively manage and use these large datasets.
Ed Black Elected President-elect of SRA
 Congratulations to our own Grants and Accounting Manager, Edward Black who has been elected President-elect of the Society of Research Administrators (SRA) International.
Congratulations to our own Grants and Accounting Manager, Edward Black who has been elected President-elect of the Society of Research Administrators (SRA) International. The SRA is dedicated to the education and professional development of research administrators working in varied organizational settings as well as the advancement of research administration as a profession around the world. As President-elect, Edward will serve as second in command of the organization and co-chair of their annual meeting. In one year, when the current President steps down, Edward will assume the office of President. After serving as President for a year Edward will serve an additional year as an advisor to the next president. Although Ed will be assuming new responsibilities in his role with the SRA, we're happy to report that he will not be leaving SCI.
See SRA Catalyst - "The SRA Western Section Election Results Are In"
CARMA Team Hosts Successful Atrial Fibrillation Symposium
 The Western Atrial Fibrillation Symposium is a comprehensive education meeting focused exclusively on atrial fibrillation. The conference features distinguished faculty and physicians from eight countries and 20 renowned clinical and research centers across the United States.
The Western Atrial Fibrillation Symposium is a comprehensive education meeting focused exclusively on atrial fibrillation. The conference features distinguished faculty and physicians from eight countries and 20 renowned clinical and research centers across the United States.The symposium has grown greatly since it's origination in 2007. This year's symposium hosted over 300 scientists from around the world. The success of the symposium is due in large part to the hard work and dedication of the University of Utah's Dr. Robert MacLeod, Dr. Nassir F. Marrouche, Dr. Mohamed H. Hamdan and the CARMA Team.
U Receives $2.9 Million for Down Syndrom Research
 University of Utah Receives $2.9 Million Grant for Groundbreaking Down Syndrome Research. Studies including Utah families will provide insights into the human brain and treatment strategies for mental disability SALT LAKE CITY – A multidisciplinary team of University of Utah investigators has received a grant for innovative research that will shed new light on the genes that cause Down syndrome (DS), as well as the defects of brain development and function that lead to intellectual disability. The $2.96 million grant co-funded by the Eunice Kennedy Shriver National Institute of Child Health and Human Development and the National Institute of Neurological Disorders and Stroke at the National Institutes of Health (NIH) will support cross-disciplinary research that studies not only genes and brain structure, but also the circuitry and chemical signals within the brain that lead to the development of DS.
University of Utah Receives $2.9 Million Grant for Groundbreaking Down Syndrome Research. Studies including Utah families will provide insights into the human brain and treatment strategies for mental disability SALT LAKE CITY – A multidisciplinary team of University of Utah investigators has received a grant for innovative research that will shed new light on the genes that cause Down syndrome (DS), as well as the defects of brain development and function that lead to intellectual disability. The $2.96 million grant co-funded by the Eunice Kennedy Shriver National Institute of Child Health and Human Development and the National Institute of Neurological Disorders and Stroke at the National Institutes of Health (NIH) will support cross-disciplinary research that studies not only genes and brain structure, but also the circuitry and chemical signals within the brain that lead to the development of DS.
The Art of Science
 February 11, 2012 - April 7, 2012
February 11, 2012 - April 7, 2012Art Talk Thursday, March 15th, 6-7pm.
Kimball Art Center, 638 Park Avenue, Park City, UT 84060
Science and art come together this month at the Kimball Art Center’s Badami Gallery. SCI Institute: The Art of Science is on display through April 7th, 2012. Some of the world’s most recognized scientists in the computing and imaging field have their ground breaking work on display.
As internationally acclaimed leaders in scientific visualization, researchers at the Scientific Computing and Imaging Institute at the University of Utah work constantly to develop new and better ways to visualize and communicate the results of scientific inquiry.
Although their visual designs are created to serve science, many of SCI’s visualizations of computer simulations of combustion or bioelectric activity in the heart and brain, or of data ranging from a galactic scale down to a cellular level, can also be viewed as works of art.
FluoRender Image Featured in Cell Picture Show
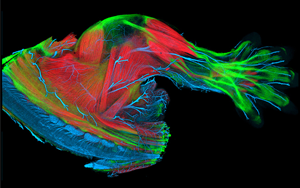 A stunning FluoRender image visualizing embryogenesis is featured in a set on cell.com's "Cell Picture Show." See "Limbs" on slide #8:
A stunning FluoRender image visualizing embryogenesis is featured in a set on cell.com's "Cell Picture Show." See "Limbs" on slide #8: Limbs
By A. Kelsey Lewis, Gabrielle Kardon, Yong Wan (University of Utah) and Ronen Schweitzer (Oregon Health and Science University)
Caption: In mice, muscles (red), tendons (green), and nerves (blue) of the limbs are largely established by ~14.5 after fertilization. In humans, this occurs after ~9 weeks of embryogenesis. Image: Here the limb of a Scleraxis-GFPmouse embryo is imaged by confocal microscopy (Nikon A1R). Scleraxis is a basic helix-turn-helix transcription factor expressed in tendon cells. Antibodies to mysosin (red) and neurofilaments (blue) label muscle and nerves, respectively. 3D rendering by FluoRender.
Utah Image Processing Group Plays Key Role in Autism Research
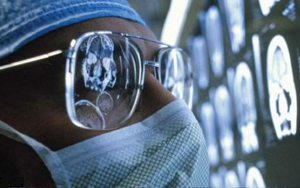 Brain Imaging Differences Evident at 6 Months in High-Risk Infants Who Later Develop Autism A new study led by the University of North Carolina at Chapel Hill found significant differences in brain development starting at age 6 months in high-risk infants who later develop autism, compared to high-risk infants who did not develop autism.
Brain Imaging Differences Evident at 6 Months in High-Risk Infants Who Later Develop Autism A new study led by the University of North Carolina at Chapel Hill found significant differences in brain development starting at age 6 months in high-risk infants who later develop autism, compared to high-risk infants who did not develop autism.The study was published online on Feb. 17 at AJP in Advance, a section of the website of the American Journal of Psychiatry. Its results are the latest from the ongoing Infant Brain Imaging Study (IBIS) Network, which is funded by the National Institutes of Health and headquartered at UNC. Piven received an NIH Autism Centers of Excellence (ACE) program network award for the IBIS Network in 2007. ACE networks consist of researchers at many facilities in locations throughout the country, all of whom work together on a single research question.
Chuck Hansen Elected IEEE Fellow
 Congratulations to Dr. Chuck Hansen who has been elected an IEEE Fellow in recognition of his extraordinary accomplishments in the development of visualization tools for large-scale scientific datasets. This honorary designation is limited to no more than one-tenth of one percent of the total voting IEEE Institute membership each year.
Congratulations to Dr. Chuck Hansen who has been elected an IEEE Fellow in recognition of his extraordinary accomplishments in the development of visualization tools for large-scale scientific datasets. This honorary designation is limited to no more than one-tenth of one percent of the total voting IEEE Institute membership each year.




