Developing the Next Generation Tools for Preoperative Planning for Implantable Cardiac Defibrillators
 BioPSE visualizing the electrical field generated by an ICD device. |
The use of ICDs has greatly increased over the last few years due to their efficacy in preventing sudden cardiac death (SCD) in patients with congenital heart defects or heart disease. These devices work by continually monitoring the rhythm of the patient's heart and immediately delivering a corrective electric shock if a life-threatening tachycardia is detected. Through this innovation, thousands of lives are saved each year. Surprisingly, these devices are sometimes implanted in newborns and older children with congenital heart defects. Pediatric patients present a particular challenge to the surgeons planning an implantation due to the wide variety of shapes and sizes of torsos. It often has proven difficult for physicians to determine the ideal placement and orientation of the electrodes prior to surgery. Accurate placement of the electrodes is crucial to ensure successful defibrillation with a minimum amount of electric current and to minimize potential damage to the heart and the surrounding tissues.
Building a Better Brain Atlas
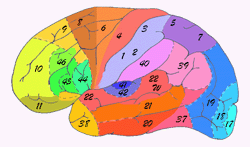 In 1909 the Broadman areas map was published which localized various functions of cortex. |
Although we've identified many structures and characteristics that are common in all human brains, in reality every brain is different and we need to improve our understanding of how brains vary between individuals. One problem that persists is that most current atlases have been based on arbitrarily chosen individuals. Even today, when intense research has been directed toward the development of digital three-dimensional atlases of the brain, most digital brain atlases so far have been based on a single subject's anatomy. This introduces a bias into the analysis when comparing individual brains to the atlas and does not provide a meaningful baseline with which to measure individual anatomical variation.
VisTrails: A New Paradigm for Dataflow Management
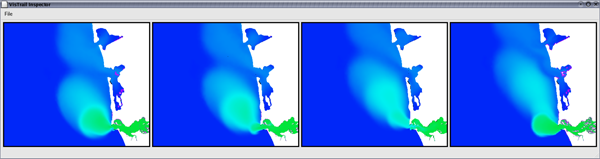 Figure 1: The VisTrails Visualization Spreadsheet. Surface salinity variation at the mouth of the Columbia River over the period of a day. The green regions represent the fresh-water discharge of the river into the ocean. A single vistrail specification is used to construct this ensemble. Each cell corresponds to a single visualization pipeline specification executed with a different timestamp value. |
The Mechanics of Angiogenesis
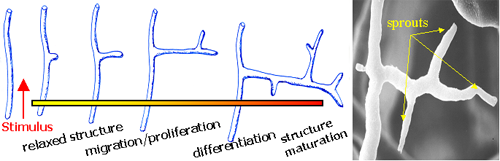 Growth of new blood vessels from existing ones. |
Embryos Exposed in 3-D
New Method Can Identify What Genes Do, Test Drugs' Safety
May 4, 2006 -- Utah and Texas researchers combined miniature medical CT scans with high-tech computer methods to produce detailed three-dimensional images of mouse embryos – an efficient new method to test the safety of medicines and learn how mutant genes cause birth defects or cancer."Our method provides a fast, high-quality and inexpensive way to visually explore the 3-D internal structure of mouse embryos so scientists can more easily and quickly see the effects of a genetic defect or chemical damage,” says Chris Johnson, a distinguished professor of computer science at the University of Utah.
A study reporting development of the new method – known as “microCT-based virtual histology” – was published recently in PLoS Genetics, an online journal of the Public Library of Science.
The study was led by Charles Keller, a pediatric cancer specialist who formerly worked as a postdoctoral fellow in the laboratory of University of Utah geneticist Mario Capecchi. Keller now is an assistant professor at the Children's Cancer Research Institute at the University of Texas Health Science Center in San Antonio.
The Center for Interactive Ray-Tracing and Photo Realistic Visualization
Almost every modern computer comes with a graphics processing unit (GPU) that implements an object-based graphics algorithm for fast 3-D graphics. The object-based algorithm in these chips was developed at the University of Utah in the 1970s. While these chips are extremely effective for video games and the visualization of moderately sized models, they cannot interactively display many of the large models that arise in computer-aided design, film animation, and scientific visualization. Researchers at the University of Utah have demonstrated that image-based ray tracing algorithms are more suited for such large-scale applications. A substantial code base has been developed in the form of two ray tracing programs. The new Center aims to improve and integrate these programs to make them appropriate for commercial use.
Top Scientific Visualization Research Problems
 Scientific visualization as currently understood and practiced is still a relatively new discipline. As a result, we visualization researchers are not necessarily accustomed to undertaking the sorts of self-examinations that other scientists routinely undergo in relation to their work. Yet if we are to create a disciplinary culture focused on matters of real scientific importance and committed to real progress, it is essential that we ask ourselves hard questions on an ongoing basis. What are the most important research issues facing us? What underlying assumptions need to be challenged and perhaps abandoned? What practices need to be reviewed? In this article, I attempt to start a discussion of these issues by proposing a list of top research problems and issues in scientific visualization. [PDF version]
Scientific visualization as currently understood and practiced is still a relatively new discipline. As a result, we visualization researchers are not necessarily accustomed to undertaking the sorts of self-examinations that other scientists routinely undergo in relation to their work. Yet if we are to create a disciplinary culture focused on matters of real scientific importance and committed to real progress, it is essential that we ask ourselves hard questions on an ongoing basis. What are the most important research issues facing us? What underlying assumptions need to be challenged and perhaps abandoned? What practices need to be reviewed? In this article, I attempt to start a discussion of these issues by proposing a list of top research problems and issues in scientific visualization. [PDF version]
CT Segmentation and 3-D Reconstruction with Morphometric Analysis for Evaluation of Occipital-Cervical Instability in Children
Down syndrome is a common chromosomal disorder that affects 0.15 percent of the total population. Individuals with Down syndrome are prone to spinal instability due to congenital abnormalities in the occipital-cervical (O-C1) joint near the base of the skull. To determine the possible abnormality causing this instability, an image-processing pipeline was created by combining several available software packages and custom made software. Patients with Down syndrome and spinal instability at the O-C1 joint were age-matched with controls. The subject data was assessed and a congenital abnormality was defined in the superior articular facets of C1 in the Down syndrome patients. The software developed helped to visualize the abnormality and could be used in a clinical setting to help aide in the diagnosis and screening for spinal instability in Down syndrome patients.
Level-Set Metamorphosis Goes Hollywood

Visualization of Vortex Flow Structures in Aerodynamics Applications
Computational Fluid Dynamics (CFD) has become an essential tool in various engineering fields. In aeronautics it is a key element in the design of modern aircrafts. The performances of today's computers combined with the increasing complexity of physical models yields numerical simulations that accurately reproduce the flow structures observed in practical experiments and permit to study their impact on flight stability. Yet, to fully exploit the huge amount of information contained in typical data sets engineers require powerful post-processing techniques that allow insight into the results of their large-scale computation.
Flow visualization aims at addressing this challenge by offering intuitive and effective depictions of interesting flow patterns. Unfortunately, many problems remain that limit the usefulness of existing methods in practical applications. Our recent work has focused on the design of new visualization techniques suitable for large-scale CFD simulations. Special emphasis was put on critical flight situations that lead to turbulent and vortical flows as well as complex and structurally involved phenomena like flow recirculation and vortex breakdown.
Visualization of Anatomic Covariance Tensor Fields
The computation, visualization, and interpretation of brain variability remains a significant challenge in computational neuroanatomy. Current deformable registration methods can generate, for each vertex of a polygonal mesh modeling the cortical surface, a distribution of displacement vectors between the individual models and their average, which can be summarized as a covariance tensor. While analysis of anatomical covariance tensor fields promises insight into the structural components of aging and disease, basic understanding of the tensor field structure is hampered by the lack of effective methods to create informative and interactive visualizations. We describe a novel application of superquadric tensor glyphs to anatomic covariance tensor fields, supplemented by colormaps of important tensor attributes. The resulting visualizations support a more detailed characterization of population variability of brain structure than possible with previous methods, while also suggesting directions for subsequent quantitative analysis.
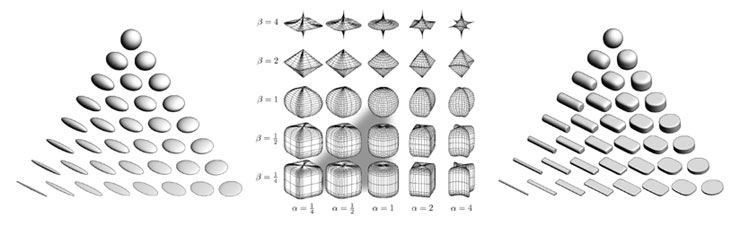 Figure 1: (a) Ellipsoid tensor glyphs, (b) Space of superquadrics, (c) Superquadric tensor glyphs. Ellipsoidal glyphs (a) suffer from visual ambiguity. The range of superquadrics (b) used for tensor glyphs is highlighted with the gray triangle. Superquadric glyphs (c) differentiate shape and convey orientation more clearly than do ellipsoids. Figure 1: (a) Ellipsoid tensor glyphs, (b) Space of superquadrics, (c) Superquadric tensor glyphs. Ellipsoidal glyphs (a) suffer from visual ambiguity. The range of superquadrics (b) used for tensor glyphs is highlighted with the gray triangle. Superquadric glyphs (c) differentiate shape and convey orientation more clearly than do ellipsoids. |
Interactive Point Based Isosurface Extraction
We have proposed a novel point-based approach to view dependent isosurface extraction. We also introduce a fast visibility query system for the view dependent traversal, which exhibits moderate memory requirements. Using this technique, we achieved an interactive interrogation of the full visible woman dataset (1GB) at more then four frames per second on a desktop computer. The point-based approach is based on an extraction scheme that classifies different sections of the isosurface into four categories. The classification is based on the size of the geometry when projected onto the screen. In particular, we use points to represent small and sub-pixel triangles, as well as large sections of the isosurface whose projection has sub-pixel size. An important issue raised by point-based processing is how to assign a normal to an isolated point representing a large, but far, section of the isosurface. We propose to define such normals during a post processing of the extracted isosurface and provide the corresponding hardware implementation.
 Figure 1: Left: A section of the visible female skeleton. Middle: A closeup view of the extracted points. Right: The final visibility mask. The color represent different levels of the mask hierarchy Figure 1: Left: A section of the visible female skeleton. Middle: A closeup view of the extracted points. Right: The final visibility mask. The color represent different levels of the mask hierarchy |
BioTensor: Visualizing Diffusion Tensor MRI
 Diffusion tensor MRI visualization is a growing field of research. The scanners are collecting better data all the time, and doctors and scientists are constantly discovering new applications for this data. However, unlike scalar and vector data, high-dimensional tensors are not always intuitive to visualize. When devising new strategies for DT-MRI visualization, it is important to understand both what exactly it is that is being measured and what insights the doctors and scientists are hoping to gain from the data.
Diffusion tensor MRI visualization is a growing field of research. The scanners are collecting better data all the time, and doctors and scientists are constantly discovering new applications for this data. However, unlike scalar and vector data, high-dimensional tensors are not always intuitive to visualize. When devising new strategies for DT-MRI visualization, it is important to understand both what exactly it is that is being measured and what insights the doctors and scientists are hoping to gain from the data.The success of diffusion magnetic resonance imaging (MRI) is deeply rooted in the powerful concept that during their random, diffusion-driven displacements, molecules probe tissue structure at a microscopic scale well beyond the usual image resolution. As diffusion is truly a three dimensional process, molecular mobility in tissues may be anisotropic, as in brain white matter. With diffusion tensor imaging (DTI), diffusion anisotropy effects can be fully extracted, characterized, and exploited, providing even more exquisite details on tissue microstructure. The most advanced application is certainly that of fiber tracking in the brain, which, in combination with functional MRI, might open a window on the important issue of connectivity. DTI has also been used to demonstrate subtle abnormalities in a variety of diseases (including stroke, multiple sclerosis, dyslexia, and schizophrenia) and is currently becoming part of many routine clinical protocols.
Scientific Visualization in Small Animal Imaging
Biomedical applications of small animal imaging are creating exciting opportunities to extend the scientific impact of visualization research. Specifically, the effective pairing of non-linear image filtering and direct volume rendering is one strategy for scientists to quickly explore and understand the volumetric scans of their specimens. Microscopic computed tomography imaging is an increasingly popular and powerful modality for small animal imaging. Here we highlight early work from collaborations at the University of Utah between the Scientific Computing and Imaging (SCI) Institute and the Department of Bioengineering, and between the SCI Institute and the Division of Pediatric Hematology-Oncology in the Department of Pediatrics. In the first instance, volume rendering provides information about the three-dimensional configuration of an electrode array implanted into the auditory nerve of a feline. In the second instance, volume rendering shows promise as a tool for visualizing bone tissue in the mouse embryo, although the signal-to-noise characteristics of the data require the use of sophisticated image pre-processing.
The data for both of these investigations was acquired with a General Electric EVS RS-9 computed tomography scanner at the University of Utah Small Animal Imaging Facility. The scanner generates 16-bit volumes roughly one gigabyte in size, with a spatial resolution of 21 x 21 x 21 microns.
Collaborative Remote Visualization
Introduction
In the last few years, scientists and researchers have given a great deal of attention to the area of remote visualization of scientific datasets within collaborative environments. This recent interest has been fueled by the use of interactive viewing as the primary means by which researchers explore large datasets. However, researchers often need to extend this interactivity in order to collaborate with colleagues who are geographically separated. Most current remote visualization tools allow multiple parties to view images from different locations, but pose problems with efficiency and user interactivity.Undergraduate Research at the SCI Institute
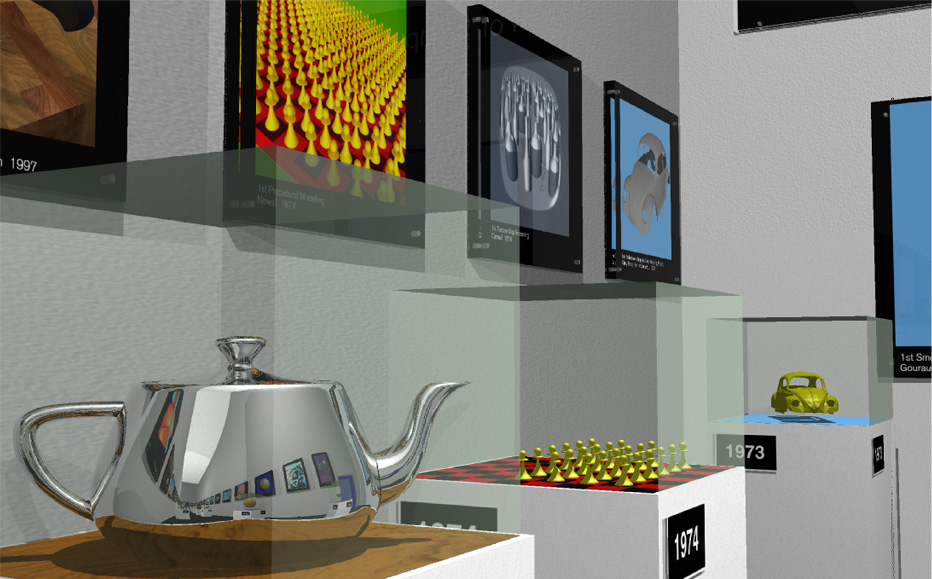
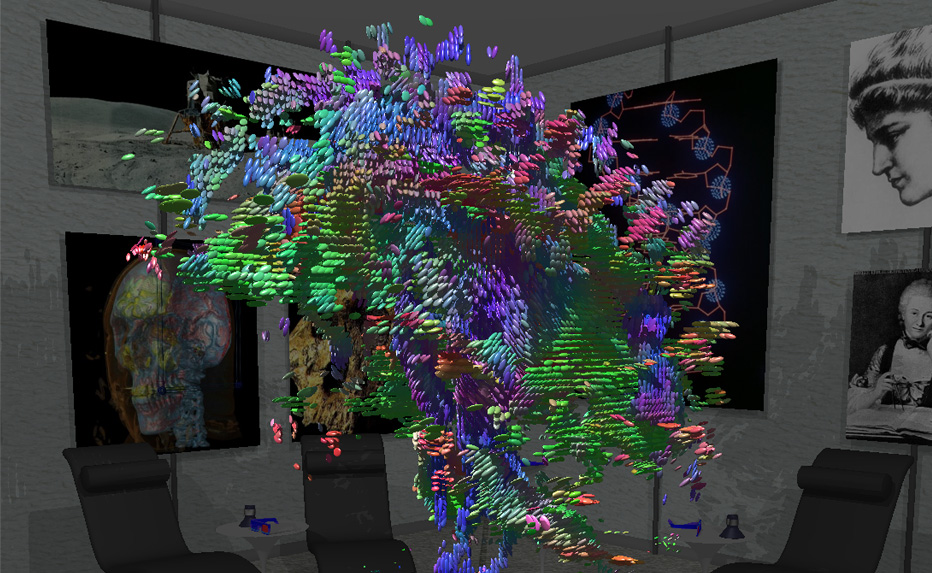
Star-Ray Interactive Ray-Tracer Debuts at SIGGRAPH 2002
 |
 |
 |
 |
 |
(left to right) Living Room Scene, Graphics Museum, Science Room, Galaxy Room, Atlantis Scene
Parametric Method for Correction of Intensity Inhomogeneity in MRI Data
Intensity inhomogeneity is one of the main obstacles for MRI data post processing. The problem requires retrospective correction due to the strong dependence of the inhomogeneity on patient anatomy and the accompanying acquisition protocol. We have developed a new method for correcting the inhomogeneities using a polynomial estimation of the bias field. The method minimizes the composite energy function to find parameters of the polynomial model. The energy function is designed to provide a robust estimation of the bias field by combining measures from histogram analysis and local gradient estimation. The method was validated on a wide range of MRI data obtained with coils of different types and under different acquisition protocols.
The developed method provides reliable estimation of the intensity inhomogeneities in MRI data. The correction times are dependant on the number of parameters the model used, the dataset size and the degree of subsampling in estimation of both local and global terms and vary from 1 to 5 minutes using a mid range PC.
Segmenting Tomographic Data
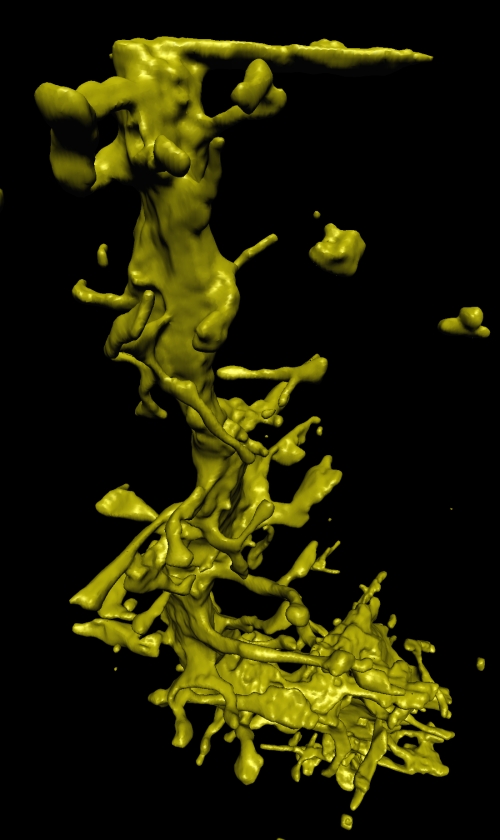 Traditionally, processing tomographic data begins with reconstructing volumes. However, when the tomographic data is incomplete, noisy, or misregistered tomographic reconstruction can produce artifacts in the volume, which makes subsequent segmentation and visualization more difficult. Researchers in the SCI institute are developing direct methods for segmenting tomographic data. The strategy is to fit 3D surface models directly to the tomographic projects, rather than the volume reconstructions. In this way, the surface fitting is not influenced by reconstruction artifacts. Implementing this strategy requires several technical advances. First is a mathematical formulation that relates object shape directly to tomographic projections. This results in a description of how surfaces should deform in order to match the tomographic data. The second advance is the use of a surface modeling technology that can accommodate a wide variety of shapes and support incremental deformations. This is done using 3D level-set models, which results in a 3D partial differential equation (PDE). The final advance is development of computational schemes that allow us to solve these PDE's efficiently. For these we have developed the incremental projection method which significantly reduces the amount of computation needed to deform these 3D surface models.
Traditionally, processing tomographic data begins with reconstructing volumes. However, when the tomographic data is incomplete, noisy, or misregistered tomographic reconstruction can produce artifacts in the volume, which makes subsequent segmentation and visualization more difficult. Researchers in the SCI institute are developing direct methods for segmenting tomographic data. The strategy is to fit 3D surface models directly to the tomographic projects, rather than the volume reconstructions. In this way, the surface fitting is not influenced by reconstruction artifacts. Implementing this strategy requires several technical advances. First is a mathematical formulation that relates object shape directly to tomographic projections. This results in a description of how surfaces should deform in order to match the tomographic data. The second advance is the use of a surface modeling technology that can accommodate a wide variety of shapes and support incremental deformations. This is done using 3D level-set models, which results in a 3D partial differential equation (PDE). The final advance is development of computational schemes that allow us to solve these PDE's efficiently. For these we have developed the incremental projection method which significantly reduces the amount of computation needed to deform these 3D surface models.
Immersive Visualization: A Research Update
What is Immersive Visualization?
The goal of visualization is to aid in the understanding of complex scientific data, typically using techniques from the fields of computer graphics and animation. To gain additional insight, immersive visualization places the user directly within the data space through virtual reality technology. The resulting immersive experience allows exploration from a first-person perspective, in contrast to the third-person interaction typical of desktop environments.The feeling of actually "being there" in a virtual environment, also known as presence, is created by fooling one or more of the user's senses with computer generated cues. In a typical system, stereo images provide a sense of visual depth and natural interaction is achieved with tracking sensors and 3D input devices. More advanced systems may also include force-feedback, spatialized audio, and/or voice recognition capabilities to increase the sense of presence.
Imagine directly navigating through a scientific dataset, much like one experiences the real world. How would it feel to investigate the interesting features of the data with your sense of touch? Would this capability be useful or not? These are the types of questions Virtual Reality (VR) researchers at the SCI institute currently seek to answer.




