Visualization
Visualization, sometimes referred to as visual data analysis, uses the graphical representation of data as a means of gaining understanding and insight into the data. Visualization research at SCI has focused on applications spanning computational fluid dynamics, medical imaging and analysis, biomedical data analysis, healthcare data analysis, weather data analysis, poetry, network and graph analysis, financial data analysis, etc.Research involves novel algorithm and technique development to building tools and systems that assist in the comprehension of massive amounts of (scientific) data. We also research the process of creating successful visualizations.
We strongly believe in the role of interactivity in visual data analysis. Therefore, much of our research is concerned with creating visualizations that are intuitive to interact with and also render at interactive rates.
Visualization at SCI includes the academic subfields of Scientific Visualization, Information Visualization and Visual Analytics.

Mike Kirby
Uncertainty Visualization
Alex Lex
Information VisualizationCenters and Labs:
- Visualization Design Lab (VDL)
- CEDMAV
- POWDER Display Wall
- Modeling, Display, and Understanding Uncertainty in Simulations for Policy Decision Making
- Topological Data Analysis for Large Network Visualization
Funded Research Projects:
Publications in Visualization:
 Topological and Statistical Methods for Complex Data, Subtitled “Tackling Large-Scale, High-Dimensional, and Multivariate Data Spaces,” J. Bennett, F. Vivodtzev, V. Pascucci (Eds.). Mathematics and Visualization, Springer Berlin Heidelberg, 2015. ISBN: 978-3-662-44899-1 This book contains papers presented at the Workshop on the Analysis of Large-scale, |
  Guided visual exploration of genomic stratifications in cancer M. Streit, A. Lex, S. Gratzl, C. Partl, D. Schmalstieg, H. Pfister, P. J. Park,, N. Gehlenborg. In Nature Methods, Vol. 11, No. 9, pp. 884--885. Sep, 2014. ISSN: 1548-7091 DOI: 10.1038/nmeth.3088 |
  ConTour: Data-Driven Exploration of Multi-Relational Datasets for Drug Discovery Christian Partl, Alexander Lex, Marc Streit, Hendrik Strobelt, Anne-Mai Wasserman, Hanspeter Pfister,, Dieter Schmalstieg. In IEEE Transactions on Visualization and Computer Graphics (VAST '14), Vol. 20, No. 12, pp. 1883--1892. 2014. ISSN: 1077-2626 DOI: 10.1109/TVCG.2014.2346752 Large scale data analysis is nowadays a crucial part of drug discovery. Biologists and chemists need to quickly explore and evaluate potentially effective yet safe compounds based on many datasets that are in relationship with each other. However, there is a lack of tools that support them in these processes. To remedy this, we developed ConTour, an interactive visual analytics technique that enables the exploration of these complex, multi-relational datasets. At its core ConTour lists all items of each dataset in a column. Relationships between the columns are revealed through interaction: selecting one or multiple items in one column highlights and re-sorts the items in other columns. Filters based on relationships enable drilling down into the large data space. To identify interesting items in the first place, ConTour employs advanced sorting strategies, including strategies based on connectivity strength and uniqueness, as well as sorting based on item attributes. ConTour also introduces interactive nesting of columns, a powerful method to show the related items of a child column for each item in the parent column. Within the columns, ConTour shows rich attribute data about the items as well as information about the connection strengths to other datasets. Finally, ConTour provides a number of detail views, which can show items from multiple datasets and their associated data at the same time. We demonstrate the utility of our system in case studies conducted with a team of chemical biologists, who investigate the effects of chemical compounds on cells and need to understand the underlying mechanisms. |
  Domino: Extracting, Comparing, and Manipulating Subsets across Multiple Tabular Datasets S. Gratzl, N. Gehlenborg, A. Lex, H. Pfister, M. Streit. In IEEE Transactions on Visualization and Computer Graphics (InfoVis '14), Vol. 20, No. 12, pp. 2023--2032. 2014. ISSN: 1077-2626 DOI: 10.1109/TVCG.2014.2346260 Answering questions about complex issues often requires analysts to take into account information contained in multiple interconnected datasets. A common strategy in analyzing and visualizing large and heterogeneous data is dividing it into meaningful subsets. Interesting subsets can then be selected and the associated data and the relationships between the subsets visualized. However, neither the extraction and manipulation nor the comparison of subsets is well supported by state-of-the-art techniques. In this paper we present Domino, a novel multiform visualization technique for effectively representing subsets and the relationships between them. By providing comprehensive tools to arrange, combine, and extract subsets, Domino allows users to create both common visualization techniques and advanced visualizations tailored to specific use cases. In addition to the novel technique, we present an implementation that enables analysts to manage the wide range of options that our approach offers. Innovative interactive features such as placeholders and live previews support rapid creation of complex analysis setups. We introduce the technique and the implementation using a simple example and demonstrate scalability and effectiveness in a use case from the field of cancer genomics. |
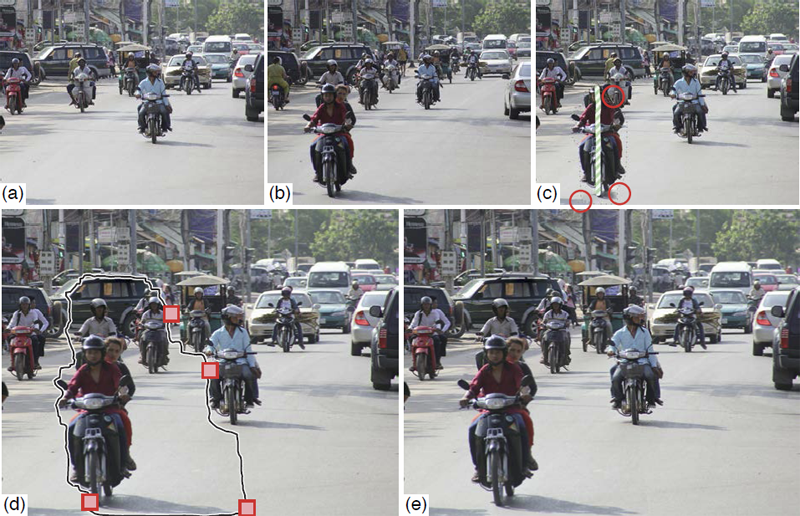
  Show me the Invisible: Visualizing Hidden Content T. Geymayer, M. Steinberger, A. Lex, M. Streit,, D. Schmalstieg. In Proceedings of the ACM SIGCHI Conference on Human Factors in Computing Systems (CHI '14), CHI '14, ACM, pp. 3705--3714. 2014. ISBN: 978-1-4503-2473-1 DOI: 10.1145/2556288.2557032 Content on computer screens is often inaccessible to users because it is hidden, e.g., occluded by other windows, outside the viewport, or overlooked. In search tasks, the efficient retrieval of sought content is important. Current software, however, only provides limited support to visualize hidden occurrences and rarely supports search synchronization crossing application boundaries. To remedy this situation, we introduce two novel visualization methods to guide users to hidden content. Our first method generates awareness for occluded or out-of-viewport content using see-through visualization. For content that is either outside the screen's viewport or for data sources not opened at all, our second method shows off-screen indicators and an on-demand smart preview. To reduce the chances of overlooking content, we use visual links, i.e., visible edges, to connect the visible content or the visible representations of the hidden content. We show the validity of our methods in a user study, which demonstrates that our technique enables a faster localization of hidden content compared to traditional search functionality and thereby assists users in information retrieval tasks. |
  Characterizing Cancer Subtypes using Dual Analysis in Caleydo C. Turkay, A. Lex, M. Streit, H. Pfister,, H. Hauser. In IEEE Computer Graphics and Applications, Vol. 34, No. 2, pp. 38--47. March, 2014. ISSN: 0272-1716 DOI: 10.1109/MCG.2014.1 Dual analysis uses statistics to describe both the dimensions and rows of a high-dimensional dataset. Researchers have integrated it into StratomeX, a Caleydo view for cancer subtype analysis. In addition, significant-difference plots show the elements of a candidate subtype that differ significantly from other subtypes, thus letting analysts characterize subtypes. Analysts can also investigate how data samples relate to their assigned subtype and other groups. This approach lets them create well-defined subtypes based on statistical properties. Three case studies demonstrate the approach's utility, showing how it reproduced findings from a published subtype characterization. |
  Mu-8: Visualizing Differences between Proteins and their Families J. Mercer, B. Pandian, A. Lex, N. Bonneel,, H. Pfister. In BMC Proceedings, Vol. 8, No. Suppl 2, pp. S5. Aug, 2014. ISSN: 1753-6561 DOI: 10.1186/1753-6561-8-S2-S5 A complete understanding of the relationship between the amino acid sequence and resulting protein function remains an open problem in the biophysical sciences. Current approaches often rely on diagnosing functionally relevant mutations by determining whether an amino acid frequently occurs at a specific position within the protein family. However, these methods do not account for the biophysical properties and the 3D structure of the protein. We have developed an interactive visualization technique, Mu-8, that provides researchers with a holistic view of the differences of a selected protein with respect to a family of homologous proteins. Mu-8 helps to identify areas of the protein that exhibit: (1) significantly different bio-chemical characteristics, (2) relative conservation in the family, and (3) proximity to other regions that have suspect behavior in the folded protein. |
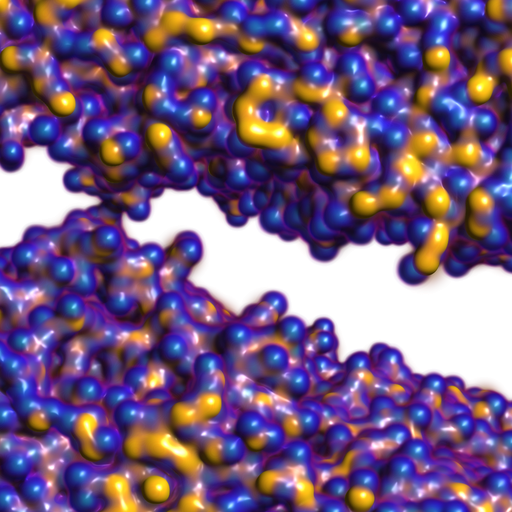
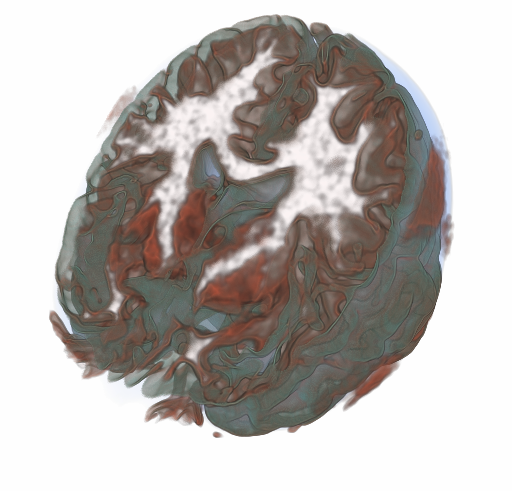
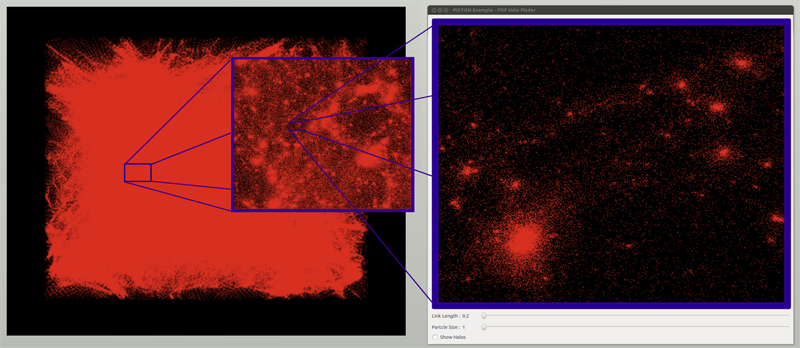
  RBF Volume Ray Casting on Multicore and Manycore CPUs A. Knoll, I. Wald, P. Navratil, A. Bowen, K. Reda, M. E. Papka, K. Gaither. In Computer Graphics Forum, Vol. 33, No. 3, Edited by H. Carr and P. Rheingans and H. Schumann, Wiley-Blackwell, pp. 71--80. June, 2014. DOI: 10.1111/cgf.12363 Modern supercomputers enable increasingly large N-body simulations using unstructured point data. The structures implied by these points can be reconstructed implicitly. Direct volume rendering of radial basis function (RBF) kernels in domain-space offers flexible classification and robust feature reconstruction, but achieving performant RBF volume rendering remains a challenge for existing methods on both CPUs and accelerators. In this paper, we present a fast CPU method for direct volume rendering of particle data with RBF kernels. We propose a novel two-pass algorithm: first sampling the RBF field using coherent bounding hierarchy traversal, then subsequently integrating samples along ray segments. Our approach performs interactively for a range of data sets from molecular dynamics and astrophysics up to 82 million particles. It does not rely on level of detail or subsampling, and offers better reconstruction quality than structured volume rendering of the same data, exhibiting comparable performance and requiring no additional preprocessing or memory footprint other than the BVH. Lastly, our technique enables multi-field, multi-material classification of particle data, providing better insight and analysis. |
  Overview and State-of-the-Art of Uncertainty Visualization G.P. Bonneau, H.C. Hege, C.R. Johnson, M.M. Oliveira, K. Potter, P. Rheingans, T. Schultz. In Scientific Visualization: Uncertainty, Multifield, Biomedical, and Scalable Visualization, Edited by M. Chen and H. Hagen and C.D. Hansen and C.R. Johnson and A. Kauffman, Springer-Verlag, pp. 3--27. 2014. ISBN: 978-1-4471-6496-8 ISSN: 1612-3786 DOI: 10.1007/978-1-4471-6497-5_1 The goal of visualization is to effectively and accurately communicate data. Visualization research has often overlooked the errors and uncertainty which accompany the scientific process and describe key characteristics used to fully understand the data. The lack of these representations can be attributed, in part, to the inherent difficulty in defining, characterizing, and controlling this uncertainty, and in part, to the difficulty in including additional visual metaphors in a well designed, potent display. However, the exclusion of this information cripples the use of visualization as a decision making tool due to the fact that the display is no longer a true representation of the data. This systematic omission of uncertainty commands fundamental research within the visualization community to address, integrate, and expect uncertainty information. In this chapter, we outline sources and models of uncertainty, give an overview of the state-of-the-art, provide general guidelines, outline small exemplary applications, and finally, discuss open problems in uncertainty visualization. |
  In-situ feature extraction of large scale combustion simulations using segmented merge trees A.G. Landge, V. Pascucci, A. Gyulassy, J.C. Bennett, H. Kolla, J. Chen, P.-T. Bremer. In Proceedings of the International Conference for High Performance Computing, Networking, Storage and Analysis (SC 2014), New Orleans, Louisana, IEEE Press, Piscataway, NJ, USA pp. 1020--1031. 2014. ISBN: 978-1-4799-5500-8 DOI: 10.1109/SC.2014.88 The ever increasing amount of data generated by scientific simulations coupled with system I/O constraints are fueling a need for in-situ analysis techniques. Of particular interest are approaches that produce reduced data representations while maintaining the ability to redefine, extract, and study features in a post-process to obtain scientific insights. |
  Efficient I/O and storage of adaptive-resolution data S. Kumar, J. Edwards, P.-T. Bremer, A. Knoll, C. Christensen, V. Vishwanath, P. Carns, J.A. Schmidt, V. Pascucci. In Proceedings of the International Conference for High Performance Computing, Networking, Storage and Analysis, IEEE Press, pp. 413--423. 2014. DOI: 10.1109/SC.2014.39 We present an efficient, flexible, adaptive-resolution I/O framework that is suitable for both uniform and Adaptive Mesh Refinement (AMR) simulations. In an AMR setting, current solutions typically represent each resolution level as an independent grid which often results in inefficient storage and performance. Our technique coalesces domain data into a unified, multiresolution representation with fast, spatially aggregated I/O. Furthermore, our framework easily extends to importance-driven storage of uniform grids, for example, by storing regions of interest at full resolution and nonessential regions at lower resolution for visualization or analysis. Our framework, which is an extension of the PIDX framework, achieves state of the art disk usage and I/O performance regardless of resolution of the data, regions of interest, and the number of processes that generated the data. We demonstrate the scalability and efficiency of our framework using the Uintah and S3D large-scale combustion codes on the Mira and Edison supercomputers. |
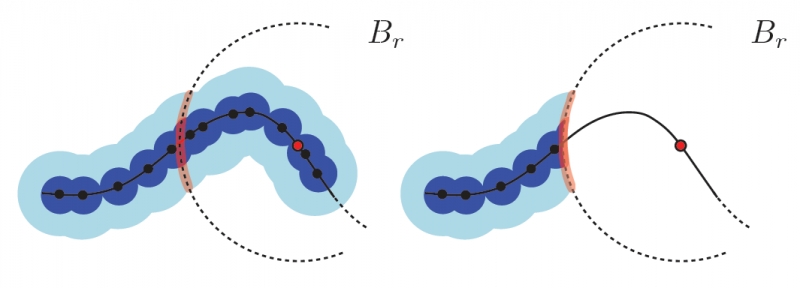
  Robust Detection of Singularities in Vector Fields H. Bhatia, A. Gyulassy, H. Wang, P.-T. Bremer, V. Pascucci . In Topological Methods in Data Analysis and Visualization III, Mathematics and Visualization, Springer International Publishing, pp. 3--18. March, 2014. DOI: 10.1007/978-3-319-04099-8_1 Recent advances in computational science enable the creation of massive datasets of ever increasing resolution and complexity. Dealing effectively with such data requires new analysis techniques that are provably robust and that generate reproducible results on any machine. In this context, combinatorial methods become particularly attractive, as they are not sensitive to numerical instabilities or the details of a particular implementation. We introduce a robust method for detecting singularities in vector fields. We establish, in combinatorial terms, necessary and sufficient conditions for the existence of a critical point in a cell of a simplicial mesh for a large class of interpolation functions. These conditions are entirely local and lead to a provably consistent and practical algorithm to identify cells containing singularities. |
 Scientific Visualization: Uncertainty, Multifield, Biomedical, and Scalable Visualization, C.D. Hansen, M. Chen, C.R. Johnson, A.E. Kaufman, H. Hagen (Eds.). Mathematics and Visualization, Springer, 2014. ISBN: 978-1-4471-6496-8 |